Ligand design for the improvement of stability of metal complex.protein hybrids.
Yokoi, N., Ueno, T., Unno, M., Matsui, T., Ikeda-Saito, M., Watanabe, Y.(2008) Chem Commun (Camb) : 229-231
- PubMed: 18092096 Search on PubMed
- DOI: https://doi.org/10.1039/b713468a
- Primary Citation Related Structures:
2Z68 - PubMed Abstract:
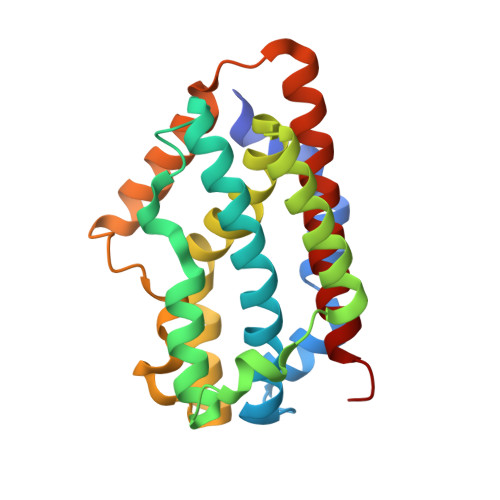
We have succeeded in improving the stability of Fe(Schiff-base).heme oxygenase (HO) hybrids by ligand design based on the crystal structure of Fe(N,N'-bis(salicylidene)-3.4-diaminobenzene propionic acid).HO.
- Department of Chemistry, Graduate School of Science, Nagoya University, Furo-cho, Chikusa-ku, Nagoya, 464-8602, Japan.
Organizational Affiliation: