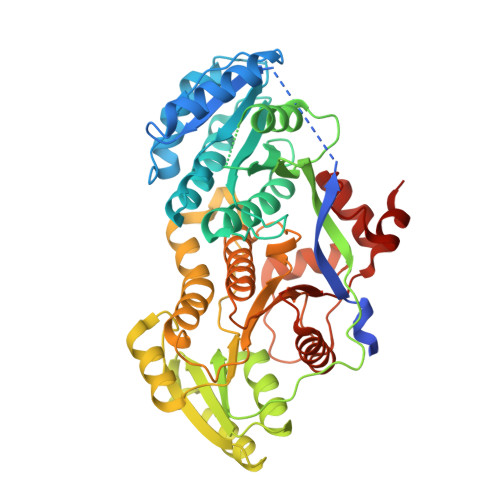
A tylosin ketoreductase reveals how chirality is determined in polyketides
Keatinge-Clay, A.T.(2007) Chem Biol 14: 898-908
- PubMed: 17719489 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2007.07.009
- Primary Citation Related Structures:
2Z5L - PubMed Abstract:
Because it controls the majority of polyketide stereocenters, the ketoreductase (KR) is a central target in engineering polyketide synthases (PKSs). To elucidate the mechanisms of stereocontrol, the structure of KR from the first module of the tylosin PKS was determined. A comparison with a recently solved erythromycin KR that operates on the same substrate explains why their products have opposite alpha-substituent chiralities. The structure reveals how polyketides are guided into the active site by key residues in different KR types. There are four types of reductase-competent KRs, each capable of fixing a unique combination of alpha-substituent and beta-hydroxyl group chiralities, as well as two types of reductase-incompetent KRs that control alpha-substituent chirality alone. A protocol to assign how a module will enforce substituent chirality based on its sequence is presented.
- Department of Biochemistry and Biophysics, University of California, San Francisco, San Francisco, CA 94158, USA. adriankc@msg.ucsf.edu
Organizational Affiliation: