Water-mediated interactions between DNA and PhoB DNA-binding/transactivation domain: NMR-restrained molecular dynamics in explicit water environment.
Yamane, T., Okamura, H., Ikeguchi, M., Nishimura, Y., Kidera, A.(2008) Proteins 71: 1970-1983
- PubMed: 18186481 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21874
- Primary Citation Related Structures:
2Z33 - PubMed Abstract:
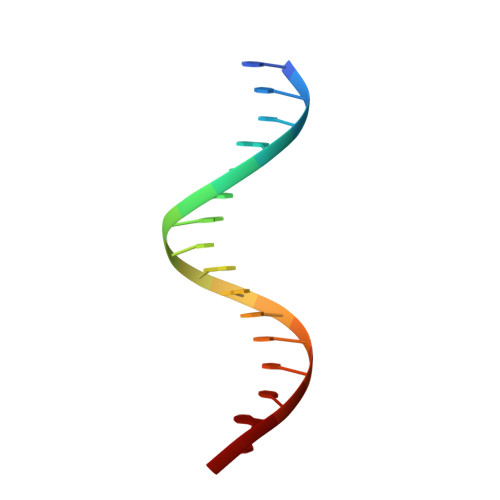
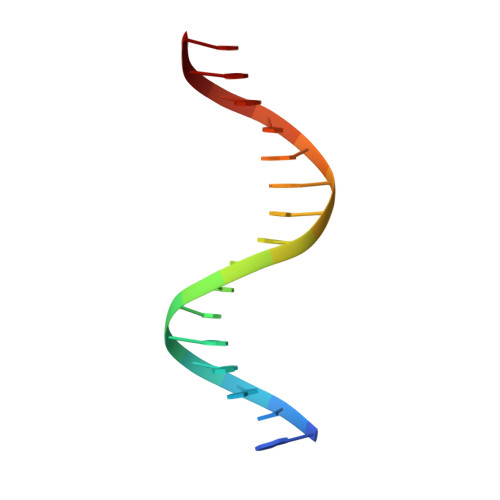
The solution structure of the complex between the transcription factor PhoB DNA-binding/transactivation domain and DNA was determined by NMR spectroscopy and simulated annealing in a periodic boundary box of explicit water with the particle mesh Ewald method. The refined structures provided better convergence and better local geometry compared with the structures determined in vacuum. The hydrogen bond interactions between the PhoB domain and DNA in the aqueous environment were fully formed. The complex structure was found to be very similar to the crystal structure, particularly at the PhoB-DNA interface, much more so than expected from the vacuum structure. These results indicate the importance of the proper treatment of electrostatic and hydration influences in describing protein-DNA interactions. The hydration structures observed for the refined structures contained most of the crystal waters as a subset. We observed that various water-mediated PhoB-DNA interactions contributed to the molecular recognition between PhoB and DNA.
- International Graduate School of Arts and Sciences, Yokohama City University, Yokohama, Japan.
Organizational Affiliation: