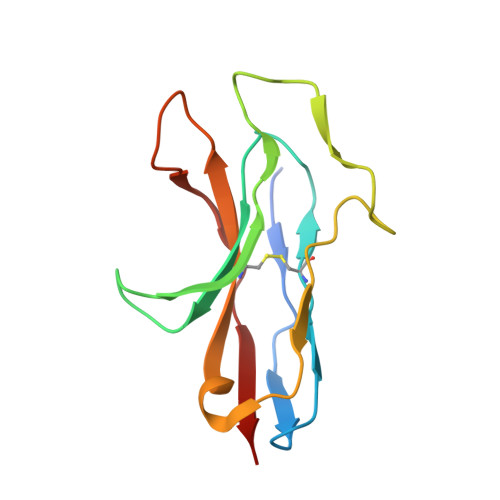
Structural insight into the specific interaction between murine SHPS-1/SIRP alpha and its ligand CD47
Nakaishi, A., Hirose, M., Yoshimura, M., Oneyama, C., Saito, K., Kuki, N., Matsuda, M., Honma, N., Ohnishi, H., Matozaki, T., Okada, M., Nakagawa, A.(2008) J Mol Biology 375: 650-660
- PubMed: 18045614 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.10.085
- Primary Citation Related Structures:
2YZ1 - PubMed Abstract:
SRC homology 2 domain-containing protein tyrosine phosphatase substrate 1 (SHPS-1 or SIRP alpha/BIT) is an immunoglobulin (Ig) superfamily transmembrane receptor and a member of the signal regulatory protein (SIRP) family involved in cell-cell interaction. SHPS-1 binds to its ligand CD47 to relay an inhibitory signal for cellular responses, whereas SIRPbeta, an activating member of the same family, does not bind to CD47 despite sharing a highly homologous ligand-binding domain with SHPS-1. To address the molecular basis for specific CD47 recognition by SHPS-1, we present the crystal structure of the ligand-binding domain of murine SHPS-1 (mSHPS-1). Folding topology revealed that mSHPS-1 adopts an I2-set Ig fold, but its overall structure resembles IgV domains of antigen receptors, although it has an extended loop structure (C'E loop), which forms a dimer interface in the crystal. Site-directed mutagenesis studies of mSHPS-1 identified critical residues for CD47 binding including sites in the C'E loop and regions corresponding to complementarity-determining regions of antigen receptors. The structural and functional features of mSHPS-1 are consistent with the human SHPS-1 structure except that human SHPS-1 has an additional beta-strand D. These results suggest that the variable complementarity-determining region-like loop structures in the binding surface of SHPS-1 are generally required for ligand recognition in a manner similar to that of antigen receptors, which may explain the diverse ligand-binding specificities of SIRP family receptors.
- Laboratory of Supramolecular Crystallography, Research Center for Structural and Functional Proteomics, Institute for Protein Research, Osaka University, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: