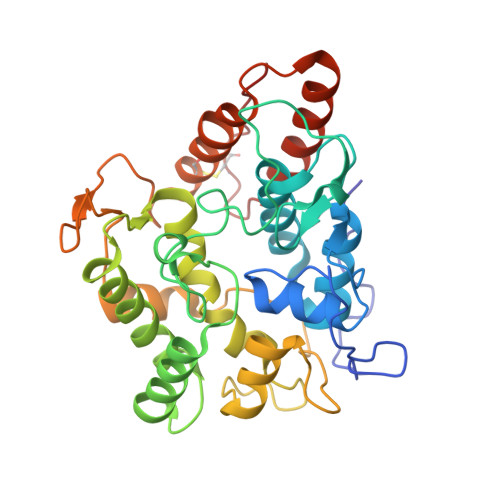
Structural Basis of Substrate Conversion in a New Aromatic Peroxygenase: P450 Functionality with Benefits
Piontek, K., Strittmatter, E., Ullrich, R., Grobe, G., Pecyna, M.J., Kluge, M., Scheibner, K., Hofrichter, M., Plattner, D.A.(2013) J Biological Chem 288: 34767
- PubMed: 24126915 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.514521
- Primary Citation Related Structures:
2YOR, 2YP1 - PubMed Abstract:
Aromatic peroxygenases (APOs) represent a unique oxidoreductase sub-subclass of heme proteins with peroxygenase and peroxidase activity and were thus recently assigned a distinct EC classification (EC 1.11.2.1). They catalyze, inter alia, oxyfunctionalization reactions of aromatic and aliphatic hydrocarbons with remarkable regio- and stereoselectivities. When compared with cytochrome P450, APOs appear to be the choice enzymes for oxyfunctionalizations in organic synthesis due to their independence from a cellular environment and their greater chemical versatility. Here, the first two crystal structures of a heavily glycosylated fungal aromatic peroxygenase (AaeAPO) are described. They reveal different pH-dependent ligand binding modes. We model the fitting of various substrates in AaeAPO, illustrating the way the enzyme oxygenates polycyclic aromatic hydrocarbons. Spatial restrictions by a phenylalanine pentad in the active-site environment govern substrate specificity in AaeAPO.
- From the Institute of Organic Chemistry, University of Freiburg, Albertstrasse 21, 79104 Freiburg.
Organizational Affiliation: