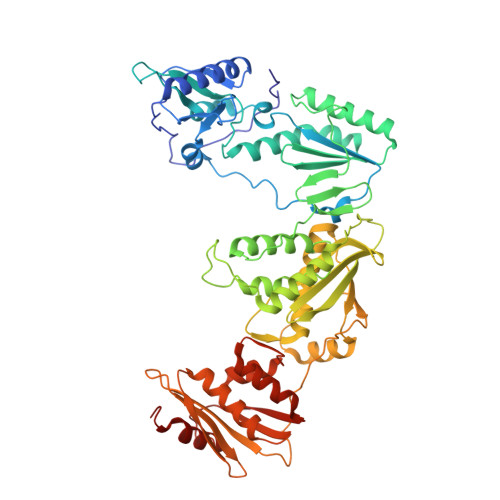
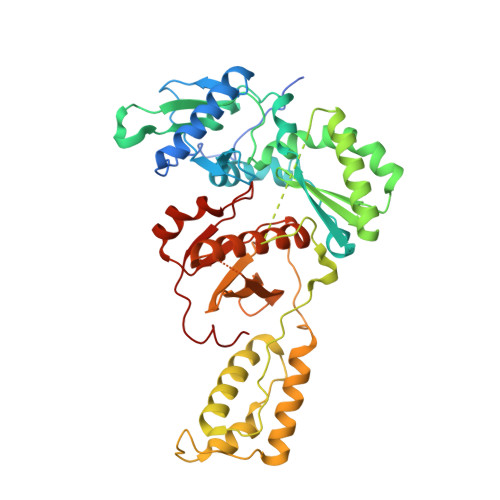
Rational Design of Potent Non-Nucleoside Inhibitors of HIV-1 Reverse Transcriptase.
Chong, P., Sebahar, P., Youngman, M., Garrido, D., Zhang, H., Stewart, E.L., Nolte, R.T., Wang, L., Ferris, R.G., Edelstein, M., Weaver, K., Mathis, A., Peat, A.(2012) J Med Chem 55: 10601
- PubMed: 23137340 Search on PubMed
- DOI: https://doi.org/10.1021/jm301294g
- Primary Citation Related Structures:
2YNF, 2YNG, 2YNH, 2YNI - PubMed Abstract:
A new series of non-nucleoside reverse transcriptase inhibitors based on an imidazole-amide biarylether scaffold has been identified and shown to possess potent antiviral activity against HIV-1, including the NNRTI-resistant Y188L mutated virus. X-ray crystallography of inhibitors bound to reverse transcriptase, including a structure of the Y188L RT protein, was used extensively to help identify and optimize the key hydrogen-bonding motif. This led directly to the design of compound 43 that exhibits remarkable antiviral activity (EC50<1 nM) against a wide range of NNRTI-resistant viruses and a favorable pharmacokinetic profile across multiple species.
- GlaxoSmithKline Research & Development, 5 Moore Drive, Research Triangle Park, North Carolina 27709, United States.
Organizational Affiliation: