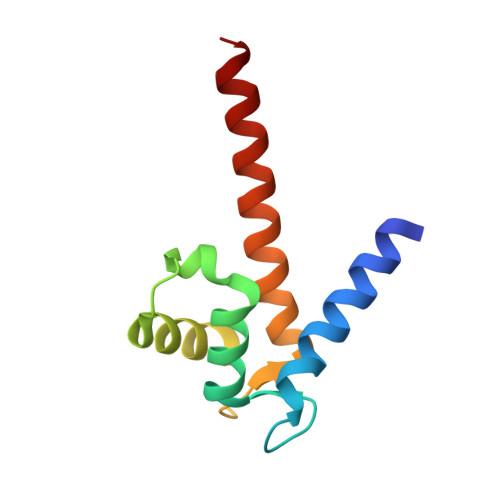
The Crystal Structure of Zebrafish S100Z: Implications for Calcium-Promoted S100 Protein Oligomerisation.
Moroz, O.V., Bronstein, I.B., Wilson, K.S.(2011) J Mol Biology 411: 1072
- PubMed: 21756915 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.06.048
- Primary Citation Related Structures:
2Y5I - PubMed Abstract:
The S100 family, with about 20 members in humans, is composed of EF-hand calcium-regulated proteins and is linked to a range of serious human diseases, including cancer and autoimmune and neurological disorders. The oldest S100 family members are found in teleosts (bony fish). The zebrafish, Danio rerio, was suggested as a promising model system for in vivo studies on S100 family functions, and we chose to investigate zebrafish S100Z as the closest homologue of the metastasis-promoting human S100A4. Here, we report the first crystal structure of an S100 protein from this organism, the calcium-bound state of S100Z to 2.03 Å resolution. Crystal packing suggests higher-order oligomerisation of S100Z dimers, with a tetramerisation interface very similar to, but even more extensive than, that reported for S100A4. The interactions are primarily through the C-terminal αIV helices from adjacent dimers in an antiparallel orientation. Structural comparisons between known S100 multimeric assemblies together with analysis of calcium-driven changes to the dimerisation cores suggest a mechanism for calcium-promoted oligomerisation of S100 proteins.
- Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York YO10 5DD, UK. olga@ysbl.york.ac.uk
Organizational Affiliation: