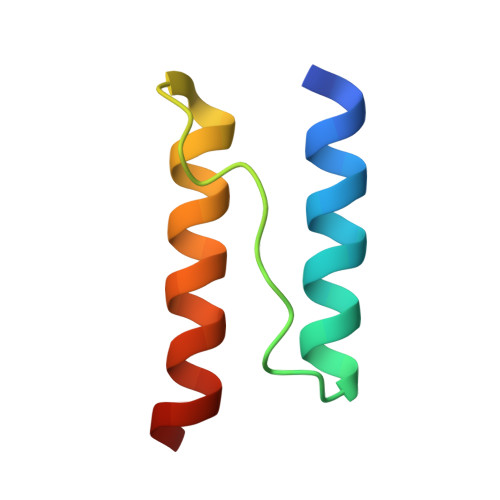
The Mechanisms of Hamp-Mediated Signaling in Transmembrane Receptors.
Ferris, H.U., Dunin-Horkawicz, S., Mondejar, L.G., Hulko, M., Hantke, K., Martin, J., Schultz, J.E., Zeth, K., Lupas, A.N., Coles, M.(2011) Structure 19: 378
- PubMed: 21397188 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2011.01.006
- Primary Citation Related Structures:
2L7H, 2L7I, 2Y0Q, 2Y0T, 2Y20, 2Y21 - PubMed Abstract:
HAMP domains mediate signal transduction in over 7500 enzyme-coupled receptors represented in all kingdoms of life. The HAMP domain of the putative archaeal receptor Af1503 has a parallel, dimeric, four-helical coiled coil structure, but with unusual core packing, related to canonical packing by concerted axial rotation of the helices. This has led to the gearbox model for signal transduction, whereby the alternate packing modes correspond to signaling states. Here we present structures of a series of Af1503 HAMP variants. We show that substitution of a conserved small side chain within the domain core (A291) for larger residues induces a gradual transition in packing mode, involving both changes in helix rotation and bundle shape, which are most prominent at the C-terminal, output end of the domain. These are correlated with activity and ligand response in vitro and in vivo by incorporating Af1503 HAMP into mycobacterial adenylyl cyclase assay systems.
- Department of Protein Evolution, Max-Planck-Institute for Developmental Biology, 72076 Tübingen, Germany.
Organizational Affiliation: