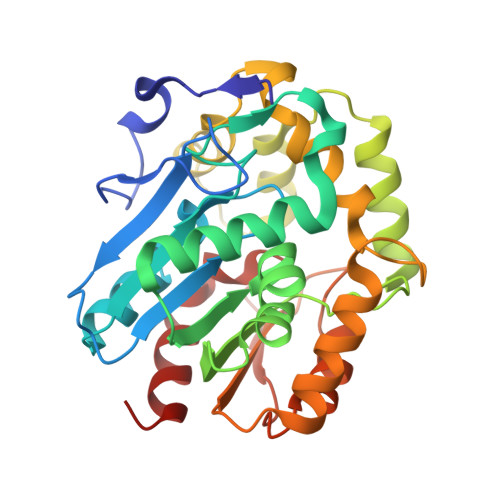
Cloning, Functional Expression, Biochemical Characterization, and Structural Analysis of a Haloalkane Dehalogenase from Plesiocystis Pacifica Sir-1.
Hesseler, M., Bogdanovic, X., Hidalgo, A., Berenguer, J., Palm, G.J., Hinrichs, W., Bornscheuer, U.T.(2011) Appl Microbiol Biotechnol 91: 1049
- PubMed: 21603934 Search on PubMed
- DOI: https://doi.org/10.1007/s00253-011-3328-x
- Primary Citation Related Structures:
2XT0 - PubMed Abstract:
A haloalkane dehalogenase (DppA) from Plesiocystis pacifica SIR-1 was identified by sequence comparison in the NCBI database, cloned, functionally expressed in Escherichia coli, purified, and biochemically characterized. The three-dimensional (3D) structure was determined by X-ray crystallography and has been refined at 1.95 Å resolution to an R-factor of 21.93%. The enzyme is composed of an α/β-hydrolase fold and a cap domain and the overall fold is similar to other known haloalkane dehalogenases. Active site residues were identified as Asp123, His278, and Asp249 and Trp124 and Trp163 as halide-stabilizing residues. DppA, like DhlA from Xanthobacter autotrophicus GJ10, is a member of the haloalkane dehalogenase subfamily HLD-I. As a consequence, these enzymes have in common the relative position of their catalytic residues within the structure and also show some similarities in the substrate specificity. The enzyme shows high preference for 1-bromobutane and does not accept chlorinated alkanes, halo acids, or halo alcohols. It is a monomeric protein with a molecular mass of 32.6 kDa and exhibits maximum activity between 33 and 37°C with a pH optimum between pH 8 and 9. The K(m) and k(cat) values for 1-bromobutane were 24.0 mM and 8.08 s(-1). Furthermore, from the 3D-structure of DppA, it was found that the enzyme possesses a large and open active site pocket. Docking experiments were performed to explain the experimentally determined substrate preferences.
- Department of Biotechnology and Enzyme Catalysis, Institute of Biochemistry, Greifswald University, Felix-Hausdorff-Str. 4, 17487 Greifswald, Germany.
Organizational Affiliation: