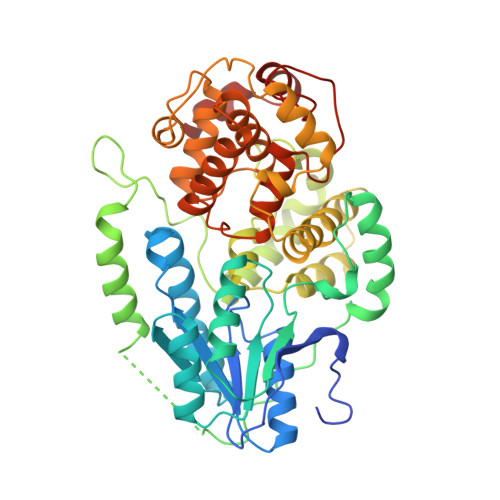
Crystal Structures of an Archaeal Class II DNA Photolyase and its Complex with Uv-Damaged Duplex DNA.
Kiontke, S., Geisselbrecht, Y., Pokorny, R., Carell, T., Batschauer, A., Essen, L.O.(2011) EMBO J 30: 4437
- PubMed: 21892138 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2011.313
- Primary Citation Related Structures:
2XRY, 2XRZ - PubMed Abstract:
Class II photolyases ubiquitously occur in plants, animals, prokaryotes and some viruses. Like the distantly related microbial class I photolyases, these enzymes repair UV-induced cyclobutane pyrimidine dimer (CPD) lesions within duplex DNA using blue/near-UV light. Methanosarcina mazei Mm0852 is a class II photolyase of the archaeal order of Methanosarcinales, and is closely related to plant and metazoan counterparts. Mm0852 catalyses light-driven DNA repair and photoreduction, but in contrast to class I enzymes lacks a high degree of binding discrimination between UV-damaged and intact duplex DNA. We solved crystal structures of Mm0852, the first one for a class II photolyase, alone and in complex with CPD lesion-containing duplex DNA. The lesion-binding mode differs from other photolyases by a larger DNA-binding site, and an unrepaired CPD lesion is found flipped into the active site and recognized by a cluster of five water molecules next to the bound 3'-thymine base. Different from other members of the photolyase-cryptochrome family, class II photolyases appear to utilize an unusual, conserved tryptophane dyad as electron transfer pathway to the catalytic FAD cofactor.
- Faculty of Chemistry, Department of Biochemistry, Philipps University, Marburg, Germany.
Organizational Affiliation: