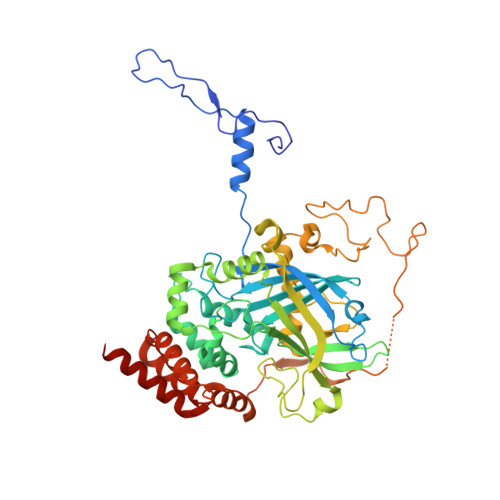
Structural Features of Peroxisomal Catalase from the Yeast Hansenula Polymorpha
Penya-Soler, E., Vega, M.C., Wilmanns, M., Williams, C.P.(2011) Acta Crystallogr D Biol Crystallogr 67: 690
- PubMed: 21795810 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444911022463
- Primary Citation Related Structures:
2XQ1 - PubMed Abstract:
The reactive oxygen species hydrogen peroxide is a byproduct of the β-oxidation process that occurs in peroxisomes. Since reactive oxygen species can cause serious damage to biomolecules, a number of scavengers control their intracellular levels. One such scavenger that is present in the peroxisome is the oxidoreductase catalase. In this study, the crystal structure of heterologously expressed peroxisomal catalase from the thermotolerant yeast Hansenula polymorpha has been determined at 2.9 Å resolution. H. polymorpha catalase is a typical peroxisomal catalase; it is tetrameric and is highly similar to catalases from other organisms. However, its hydrogen peroxide-degrading activity is higher than those of a number of other catalases for which structural data are available. Structural superimpositions indicate that the nature of the major channel, the path for hydrogen peroxide to the active site, varies from those seen in other catalase structures, an observation that may account for the high activity of H. polymorpha catalase.
- Department of Structural and Quantitative Biology, Centro de Investigaciones Biológicas (CIB-CSIC), Ramiro de Maetzu, 28040 Madrid, Spain.
Organizational Affiliation: