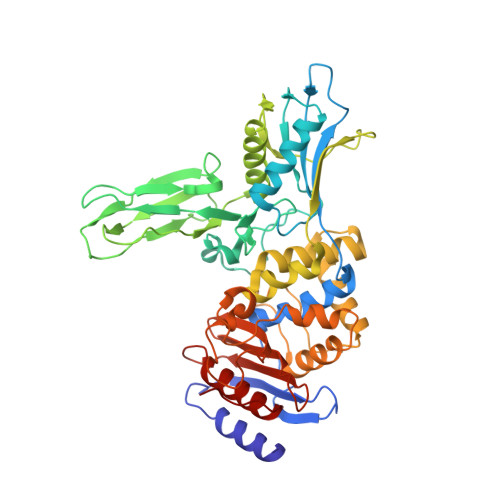
Structure Guided Development of Potent Reversibly Binding Penicillin Binding Protein Inhibitors
Woon, E.C.Y., Zervosen, A., Sauvage, E., Simmons, K.J., Ivec, M., Inglis, S.R., Fishwick, C.W.G., Gobec, S., Charlier, P., Luxen, A., Schofield, C.J.(2011) ACS Med Chem Lett 2: 219
- PubMed: 24900305 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml100260x
- Primary Citation Related Structures:
2XK1, 2XLN - PubMed Abstract:
Following from the evaluation of different types of electrophiles, combined modeling and crystallographic analyses are used to generate potent boronic acid based inhibitors of a penicillin binding protein. The results suggest that a structurally informed approach to penicillin binding protein inhibition will be useful for the development of both improved reversibly binding inhibitors, including boronic acids, and acylating inhibitors, such as β-lactams.
- Chemistry Research Laboratory, Department of Chemistry, University of Oxford , 12 Mansfield Road, Oxford OX1 3TA, United Kingdom.
Organizational Affiliation: