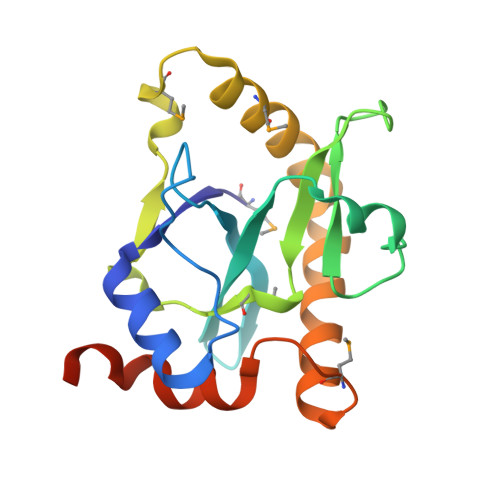
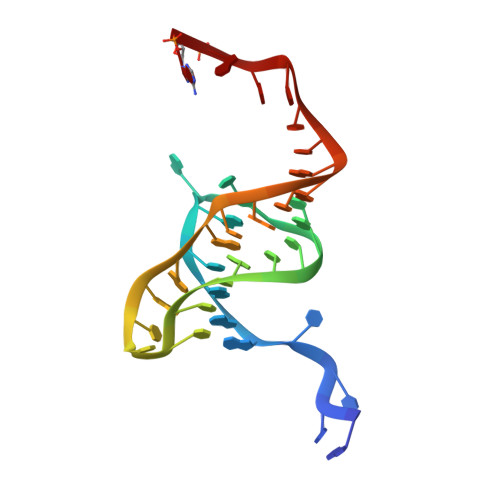
The Phage Abortive Infection System, Toxin, Functions as a Protein-RNA Toxin-Antitoxin Pair.
Fineran, P.C., Blower, T.R., Foulds, I.J., Humphreys, D.P., Lilley, K.S., Salmond, G.P.C.(2009) Proc Natl Acad Sci U S A 106: 894
- PubMed: 19124776 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0808832106
- Primary Citation Related Structures:
2XDD - PubMed Abstract:
Various mechanisms exist that enable bacteria to resist bacteriophage infection. Resistance strategies include the abortive infection (Abi) systems, which promote cell death and limit phage replication within a bacterial population. A highly effective 2-gene Abi system from the phytopathogen Erwinia carotovora subspecies atroseptica, designated ToxIN, is described. The ToxIN Abi system also functions as a toxin-antitoxin (TA) pair, with ToxN inhibiting bacterial growth and the tandemly repeated ToxI RNA antitoxin counteracting the toxicity. TA modules are currently divided into 2 classes, protein and RNA antisense. We provide evidence that ToxIN defines an entirely new TA class that functions via a novel protein-RNA mechanism, with analogous systems present in diverse bacteria. Despite the debated role of TA systems, we demonstrate that ToxIN provides viral resistance in a range of bacterial genera against multiple phages. This is the first demonstration of a novel mechanistic class of TA systems and of an Abi system functioning in different bacterial genera, both with implications for the dynamics of phage-bacterial interactions.
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1QW, United Kingdom.
Organizational Affiliation: