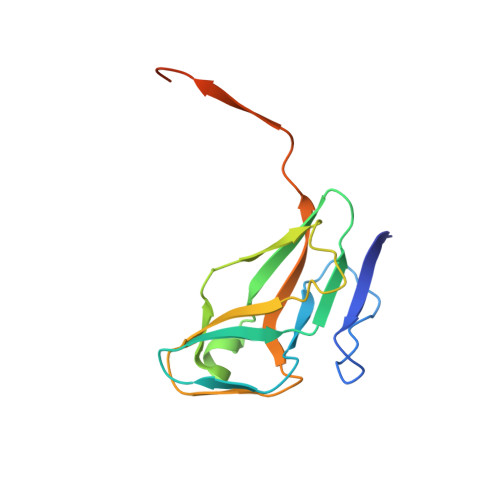
The Structure of the Genomic Bacillus Subtilis Dutpase: Novel Features in the Phe-Lid.
Garcia-Nafria, J., Burchell, L., Takezawa, M., Rzechorzek, N., Fogg, M., Wilson, K.S.(2010) Acta Crystallogr D Biol Crystallogr 66: 953
- PubMed: 20823546 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444910026272
- Primary Citation Related Structures:
2XCD, 2XCE - PubMed Abstract:
dUTPases are a ubiquitous family of enzymes that are essential for all organisms and catalyse the breakdown of 2-deoxyuridine triphosphate (dUTP). In Bacillus subtilis there are two homotrimeric dUTPases: a genomic and a prophage form. Here, the structures of the genomic dUTPase and of its complex with the substrate analogue dUpNHpp and calcium are described, both at 1.85 A resolution. The overall fold resembles that of previously solved trimeric dUTPases. The C-terminus, which contains one of the conserved sequence motifs, is disordered in both structures. The crystal of the complex contains six independent protomers which accommodate six dUpNHpp molecules, with three triphosphates in the trans conformation and the other three in the active gauche conformation. The structure of the complex confirms the role of several key residues that are involved in ligand binding and the position of the catalytic water. Asp82, which has previously been proposed to act as a general base, points away from the active site. In the complex Ser64 reorients in order to hydrogen bond the phosphate chain of the substrate. A novel feature has been identified: the position in the sequence of the ;Phe-lid', which packs against the uracil moiety, is adjacent to motif III, whereas in all other dUTPase structures the lid is in a conserved position in motif V of the flexible C-terminal arm. This requires a reconsideration of some aspects of the accepted mechanism.
- Department of Chemistry, University of York, England.
Organizational Affiliation: