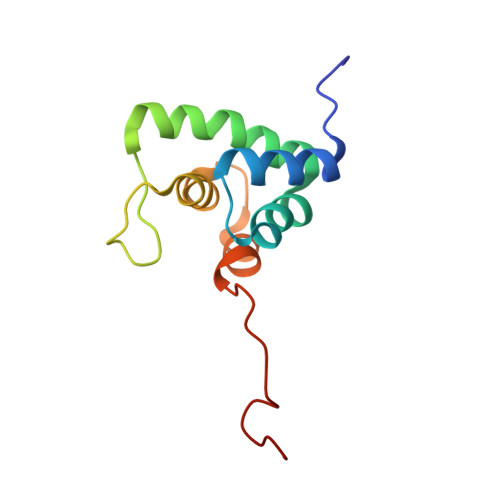
Structure and RNA Recognition by the Snrna and Snorna Transport Factor Phax.
Mourao, A., Varrot, A., Mackereth, C.D., Cusack, S., Sattler, M.(2010) RNA 16: 1205
- PubMed: 20430857 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.2009910
- Primary Citation Related Structures:
2W4S, 2XC7 - PubMed Abstract:
Small nuclear and small nucleolar RNAs (snRNAs and snoRNAs) are critical components of snRNPs and snoRNPs and play an essential role in the maturation of, respectively, mRNAs and rRNAs within the nucleus of eukaryotic cells. Complex and specific pathways exist for the assembly of snRNPs and snoRNPs, involving, for instance, nucleocytoplasmic transport of snRNAs and intranuclear transport between compartments of snoRNAs. The phosphorylated adaptor for nuclear export (PHAX) is required for nuclear export of snRNAs in metazoans and also involved in the intranuclear transport of snoRNAs to Cajal bodies. PHAX contains a conserved single-stranded nucleic acid binding domain (RNA_GG_bind domain) with no sequence homology with any other known RNA-binding module. Here, we report NMR and X-ray crystallography studies that elucidate the structural basis for RNA recognition by the PHAX RNA-binding domain (PHAX-RBD). The crystal structure of the RNA_GG_bind domain from the parasite Cryptosporidium parvum (Cp RBD) forms well-folded dimers in solution in the absence of any ligand. The human PHAX-RBD is monomeric and only adopts a tertiary fold upon RNA binding. The PHAX-RBD represents a novel helical fold and binds single-stranded RNA with micromolar affinity without sequence specificity. RNA recognition by human PHAX-RBD is consistent with mutational analysis that affects RNA binding and PHAX-mediated nuclear export. Our data suggest that the PHAX-RBD mediates auxiliary RNA contacts with the snRNA and snoRNA substrates that are required for transport and/or substrate release.
- Institute of Structural Biology, Helmholtz Zentrum München, 85764 Neuherberg, Germany.
Organizational Affiliation: