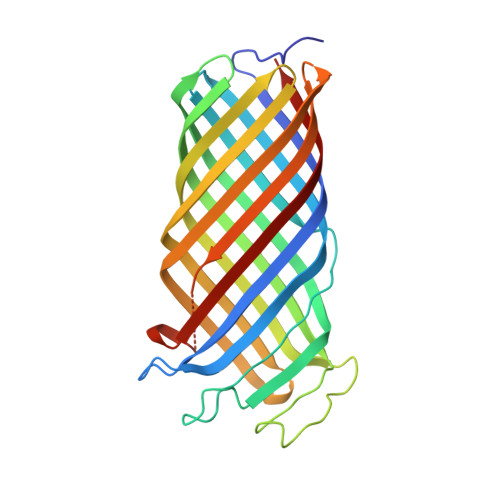
An Active Site Water Network in the Plasminogen Activator Pla from Yersinia Pestis
Eren, E., Murphy, M., Goguen, J., van den Berg, B.(2010) Structure 18: 809
- PubMed: 20637417 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2010.03.013
- Primary Citation Related Structures:
2X4M, 2X55, 2X56 - PubMed Abstract:
The plasminogen activator Pla from Yersinia pestis is an outer membrane protease (omptin) that is important for the virulence of plague. Here, we present the high-resolution crystal structure of wild-type, enzymatically active Pla at 1.9 A. The structure shows a water molecule located between active site residues D84 and H208, which likely corresponds to the nucleophilic water. A number of other water molecules are present in the active site, linking residues important for enzymatic activity. The R211 sidechain in loop L4 is close to the nucleophilic water and possibly involved in the stabilization of the oxyanion intermediate. Subtle conformational changes of H208 result from the binding of lipopolysaccharide to the outside of the barrel, explaining the unusual dependence of omptins on lipopolysaccharide for activity. The Pla structure suggests a model for the interaction with plasminogen substrate and provides a more detailed understanding of the catalytic mechanism of omptin proteases.
- Program in Molecular Medicine, University of Massachusetts Medical School, Worcester, MA 01605, USA.
Organizational Affiliation: