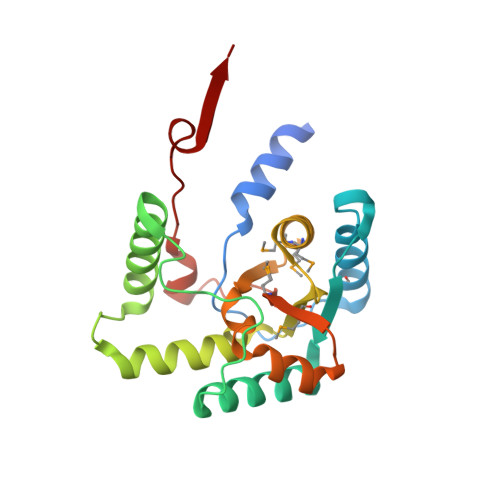
Biological and Structural Characterization of the Mycobacterium Smegmatis Nitroreductase Nfnb, and its Role in Benzothiazinone Resistance
Manina, G., Bellinzoni, M., Pasca, M.R., Neres, J., Milano, A., Ribeiro, A.L., Buroni, S., Skovierova, H., Dianiskova, P., Mikusova, K., Marak, J., Makarov, V., Giganti, D., Haouz, A., Lucarelli, A.P., Degiacomi, G., Piazza, A., Chiarelli, L.R., De Rossi, E., Salina, E., Cole, S.T., Alzari, P.M., Riccardi, G.(2010) Mol Microbiol 77: 1172
- PubMed: 20624223 Search on PubMed
- DOI: https://doi.org/10.1111/j.1365-2958.2010.07277.x
- Primary Citation Related Structures:
2WZV, 2WZW - PubMed Abstract:
Tuberculosis is still a leading cause of death in developing countries, for which there is an urgent need for new pharmacological agents. The synthesis of the novel antimycobacterial drug class of benzothiazinones (BTZs) and the identification of their cellular target as DprE1 (Rv3790), a component of the decaprenylphosphoryl-β-d-ribose 2'-epimerase complex, have been reported recently. Here, we describe the identification and characterization of a novel resistance mechanism to BTZ in Mycobacterium smegmatis. The overexpression of the nitroreductase NfnB leads to the inactivation of the drug by reduction of a critical nitro-group to an amino-group. The direct involvement of NfnB in the inactivation of the lead compound BTZ043 was demonstrated by enzymology, microbiological assays and gene knockout experiments. We also report the crystal structure of NfnB in complex with the essential cofactor flavin mononucleotide, and show that a common amino acid stretch between NfnB and DprE1 is likely to be essential for the interaction with BTZ. We performed docking analysis of NfnB-BTZ in order to understand their interaction and the mechanism of nitroreduction. Although Mycobacterium tuberculosis seems to lack nitroreductases able to inactivate these drugs, our findings are valuable for the design of new BTZ molecules, which may be more effective in vivo.
- Dipartimento di Genetica e Microbiologia, Università degli Studi di Pavia, via Ferrata, 1, 27100 Pavia, Italy.
Organizational Affiliation: