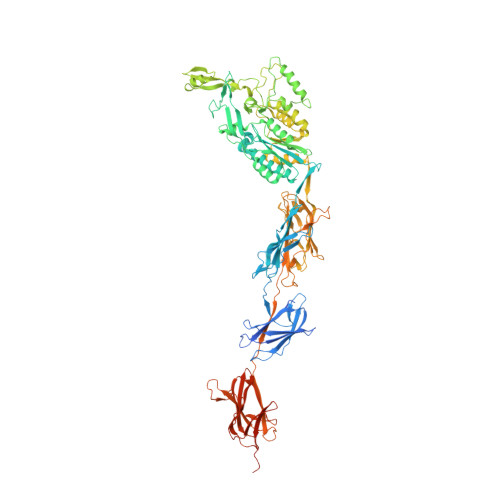
Structural Basis of Host Cell Recognition by the Pilus Adhesin from Streptococcus Pneumoniae
Izore, T., Contreras-Martel, C., El-Mortaji, L., Manzano, C., Terrasse, R., Vernet, T., Di-Guilmi, A.M., Dessen, A.(2010) Structure 18: 106
- PubMed: 20152157 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2009.10.019
- Primary Citation Related Structures:
2WW8 - PubMed Abstract:
Pili are fibrous virulence factors associated directly to the bacterial surface that play critical roles in adhesion and recognition of host cell receptors. The human pathogen Streptococcus pneumoniae carries a single pilus-related adhesin (RrgA) that is key for infection establishment and provides protection from bacterial challenge in animal infection models, but details of these roles remain unclear. Here we report the high-resolution crystal structure of RrgA, a 893-residue elongated macromolecule whose fold contains four domains presenting both eukaryotic and prokaryotic origins. RrgA harbors an integrin I collagen-recognition domain decorated with two inserted "arms" that fold into a positively charged cradle, as well as three "stalk-forming" domains. We show by site-specific mutagenesis, mass spectrometry, and thermal shift assays that intradomain isopeptide bonds play key roles in stabilizing RrgA's stalk. The high sequence similarity between RrgA and its homologs in other Gram-positive microorganisms suggests common strategies for ECM recognition and immune evasion.
- Institut de Biologie Structurale Jean-Pierre Ebel, UMR 5075 (CEA, CNRS, UJF), Grenoble, France.
Organizational Affiliation: