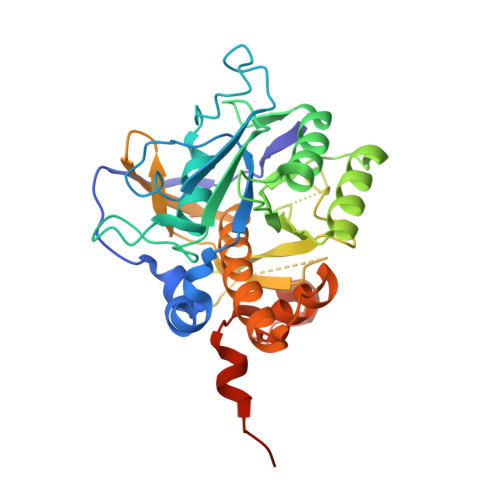
Crystal Structure of an Intracellular Subtilisin Reveals Novel Structural Features Unique to This Subtilisin Family.
Vevodova, J., Gamble, M., Kunze, G., Ariza, A., Dodson, E., Jones, D.D., Wilson, K.S.(2010) Structure 18: 744
- PubMed: 20541512 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2010.03.008
- Primary Citation Related Structures:
2WV7, 2WWT, 2X8J - PubMed Abstract:
The intracellular subtilisin proteases (ISPs) are the only known members of the important and ubiquitous subtilisin family that function exclusively within the cell, constituting a major component of the degradome in many Gram-positive bacteria. The first ISP structure reported herein at a spacing of 1.56 A reveals features unique among subtilisins that has enabled potential functional and physiological roles to be assigned to sequence elements exclusive to the ISPs. Unlike all other subtilisins, ISP from B. clausii is dimeric, with residues from the C terminus making a major contribution to the dimer interface by crossing over to contact the partner subunit. A short N-terminal extension binds back across the active site to provide a potential novel regulatory mechanism of intrinsic proteolytic activity: a proline residue conserved throughout the ISPs introduces a kink in the polypeptide backbone that lifts the target peptide bond out of reach of the catalytic residues.
- Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York YO10 5YW, UK.
Organizational Affiliation: