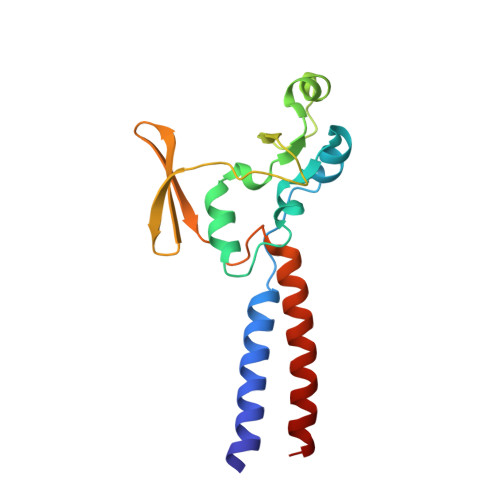
Crystal structure of the MukB hinge domain with coiled-coil stretches and its functional implications.
Ku, B., Lim, J.H., Shin, H.C., Shin, S.Y., Oh, B.H.(2010) Proteins 78: 1483-1490
- PubMed: 20034111 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22664
- Primary Citation Related Structures:
2WMM - PubMed Abstract:
The structural maintenance of chromosomes (SMC) family proteins are commonly found in the multiprotein complexes involved in chromosome organization, including chromosome condensation and sister chromatid cohesion. These proteins are characterized by forming a V-shaped homo- or heterodimeric structure with two long coiled-coil arms having two ATPase head domains at the distal ends. The hinge domain, located in the middle of the coiled coil, forms the dimer interface. In addition to being the dimerization module, SMC hinges appear to play other roles, including the gateway function for DNA entry into the cohesin complex. Herein, we report the homodimeric structure of the hinge domain of Escherichia coli MukB, which forms a prokaryotic condensin complex with two non-SMC subunits, MukE and MukF. In contrast with SMC hinge of Thermotoga maritima which has a sizable central hole at the dimer interface, MukB hinge forms a constricted dimer interface lacking a hole. Under our assay conditions, MukB hinge does not interact with DNA in accordance with the absence of a notable positively charged surface patch. The function of MukB hinge appears to be limited to dimerization of two copies of MukB molecules.
- Department of Biological Sciences, KAIST Institute for the Biocentury, Daejeon, Korea.
Organizational Affiliation: