Chimeric Microtubule Disruptors.
Leese, M.P., Jourdan, F.L., Kimberley, M.R., Cozier, G.E., Thiyagarajan, N., Stengel, C., Regis-Lydi, S., Foster, P.A., Newman, S.P., Acharya, K.R., Ferrandis, E., Purohit, A., Reed, M.J., Potter, B.V.L.(2010) Chem Commun (Camb) 46: 2907
- PubMed: 20386818 Search on PubMed
- DOI: https://doi.org/10.1039/c002558e
- Primary Citation Related Structures:
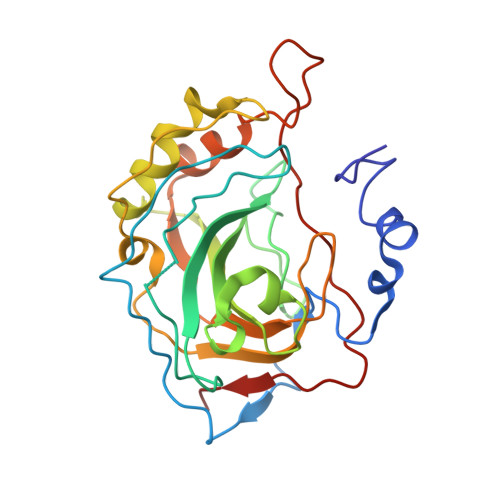
2WD2 - PubMed Abstract:
A chimeric approach is used to discover microtubule disruptors with excellent in vitro activity and oral bioavailability; a ligand-protein interaction with carbonic anhydrase that enhances bioavailability is characterised by protein X-ray crystallography. Dosing of a representative chimera in a tumour xenograft model confirms the excellent therapeutic potential of the class.
- Medicinal Chemistry & Sterix Ltd., Department of Pharmacy & Pharmacology, University of Bath, Claverton Down, Bath, BA2 7AY, UK.
Organizational Affiliation: