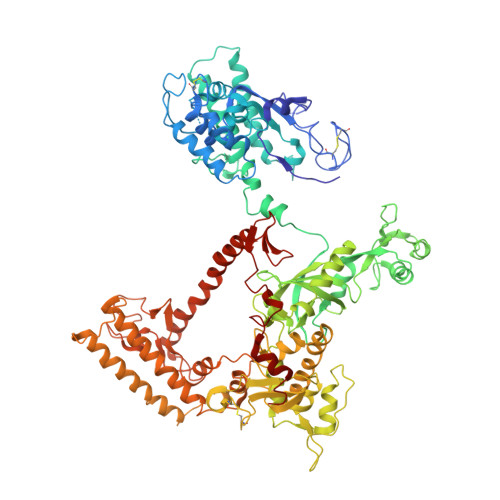
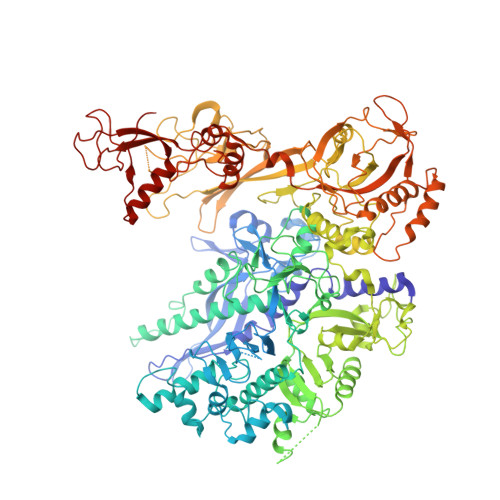
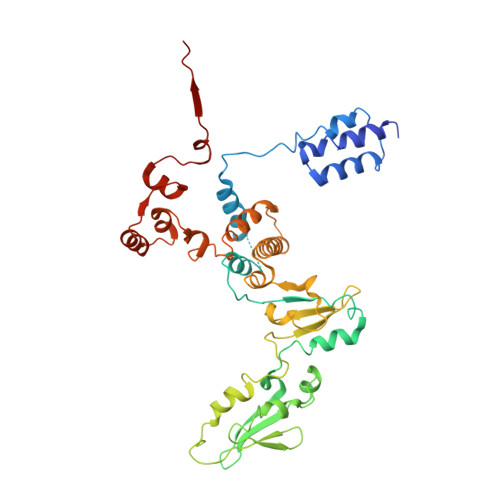
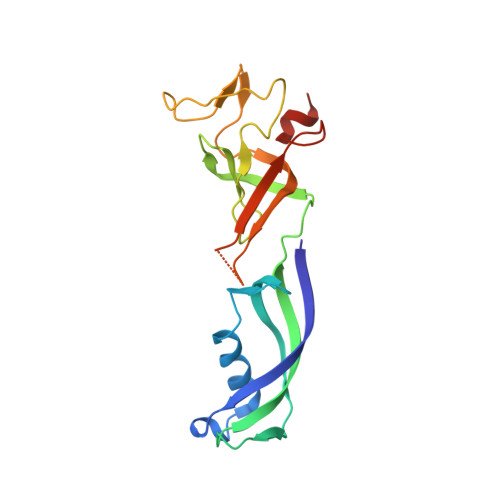
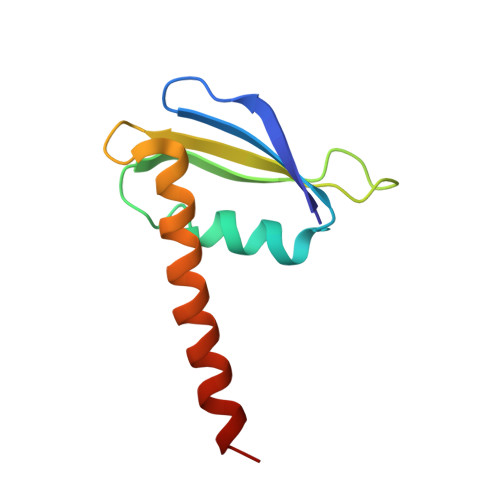
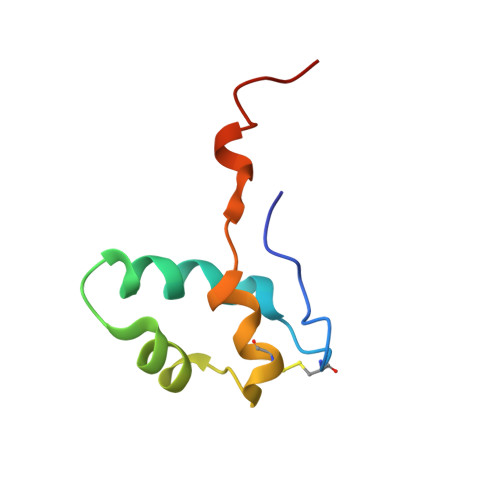
Evolution of Complex RNA Polymerase: The Complete Archaeal RNA Polymerase Structure
Korkhin, Y., Unligil, U.M., Littlefield, O., Nelson, P.J., Stuart, D.I., Sigler, P.B., Bell, S.D., Abrescia, N.G.A.(2009) PLoS Biol 7: E102
- PubMed: 19419240 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.1000102
- Primary Citation Related Structures:
2WAQ, 2WB1 - PubMed Abstract:
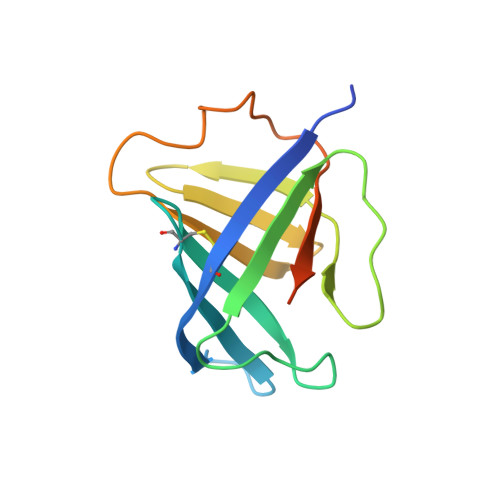
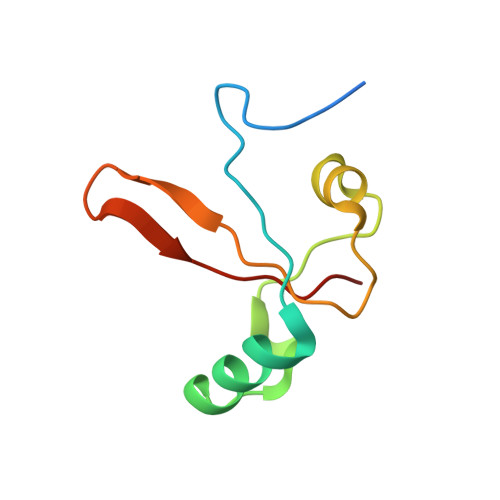
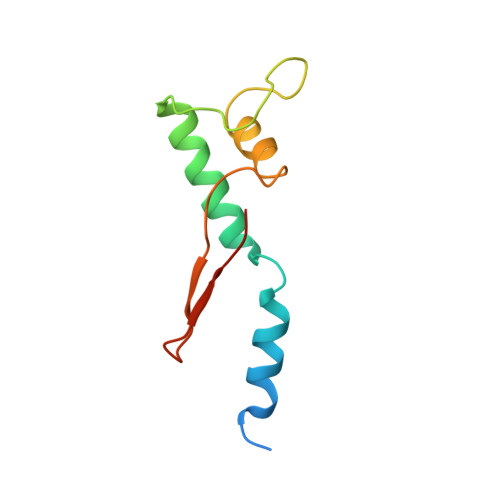
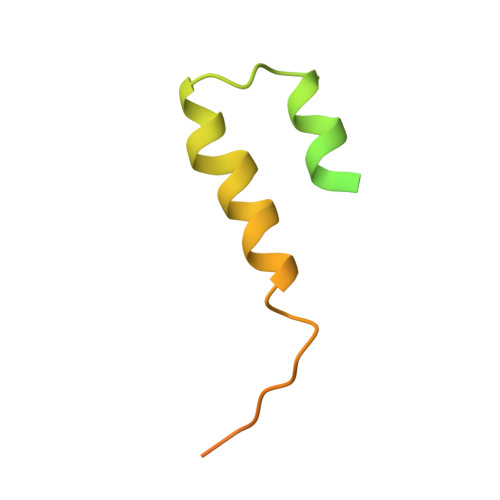
The archaeal RNA polymerase (RNAP) shares structural similarities with eukaryotic RNAP II but requires a reduced subset of general transcription factors for promoter-dependent initiation. To deepen our knowledge of cellular transcription, we have determined the structure of the 13-subunit DNA-directed RNAP from Sulfolobus shibatae at 3.35 Å resolution. The structure contains the full complement of subunits, including RpoG/Rpb8 and the equivalent of the clamp-head and jaw domains of the eukaryotic Rpb1. Furthermore, we have identified subunit Rpo13, an RNAP component in the order Sulfolobales, which contains a helix-turn-helix motif that interacts with the RpoH/Rpb5 and RpoA'/Rpb1 subunits. Its location and topology suggest a role in the formation of the transcription bubble.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut, USA.
Organizational Affiliation: