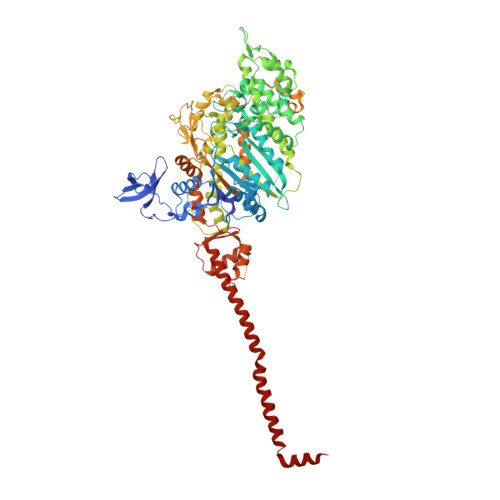
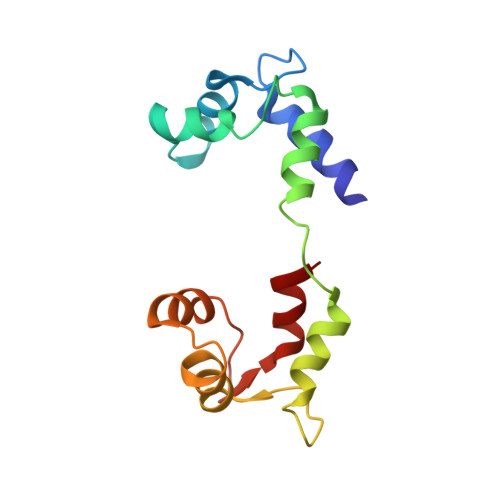
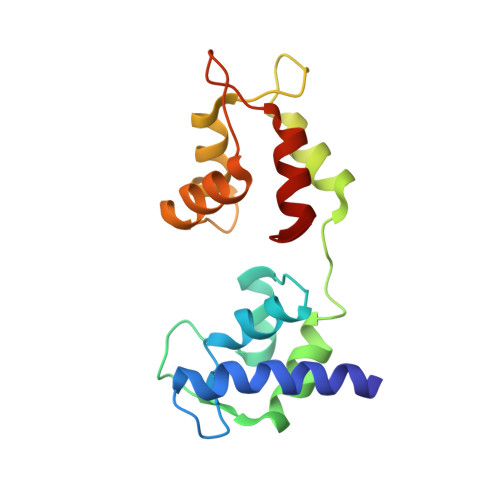
Electron Tomography of Cryofixed, Isometrically Contracting Insect Flight Muscle Reveals Novel Actin-Myosin Interactions
Wu, S., Liu, J., Reedy, M.C., Tregear, R.T., Winkler, H., Franzini-Armstrong, C., Sasaki, H., Lucaveche, C., Goldman, Y.E., Reedy, M.K., Taylor, K.A.(2010) PLoS One 5: 12643
- PubMed: 20844746 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0012643
- Primary Citation Related Structures:
2W49, 2W4A, 2W4G, 2W4T - PubMed Abstract:
Isometric muscle contraction, where force is generated without muscle shortening, is a molecular traffic jam in which the number of actin-attached motors is maximized and all states of motor action are trapped with consequently high heterogeneity. This heterogeneity is a major limitation to deciphering myosin conformational changes in situ. We used multivariate data analysis to group repeat segments in electron tomograms of isometrically contracting insect flight muscle, mechanically monitored, rapidly frozen, freeze substituted, and thin sectioned. Improved resolution reveals the helical arrangement of F-actin subunits in the thin filament enabling an atomic model to be built into the thin filament density independent of the myosin. Actin-myosin attachments can now be assigned as weak or strong by their motor domain orientation relative to actin. Myosin attachments were quantified everywhere along the thin filament including troponin. Strong binding myosin attachments are found on only four F-actin subunits, the "target zone", situated exactly midway between successive troponin complexes. They show an axial lever arm range of 77°/12.9 nm. The lever arm azimuthal range of strong binding attachments has a highly skewed, 127° range compared with X-ray crystallographic structures. Two types of weak actin attachments are described. One type, found exclusively in the target zone, appears to represent pre-working-stroke intermediates. The other, which contacts tropomyosin rather than actin, is positioned M-ward of the target zone, i.e. the position toward which thin filaments slide during shortening. We present a model for the weak to strong transition in the myosin ATPase cycle that incorporates azimuthal movements of the motor domain on actin. Stress/strain in the S2 domain may explain azimuthal lever arm changes in the strong binding attachments. The results support previous conclusions that the weak attachments preceding force generation are very different from strong binding attachments.
- Institute of Molecular Biophysics, Florida State University, Tallahassee, Florida, United States of America.
Organizational Affiliation: