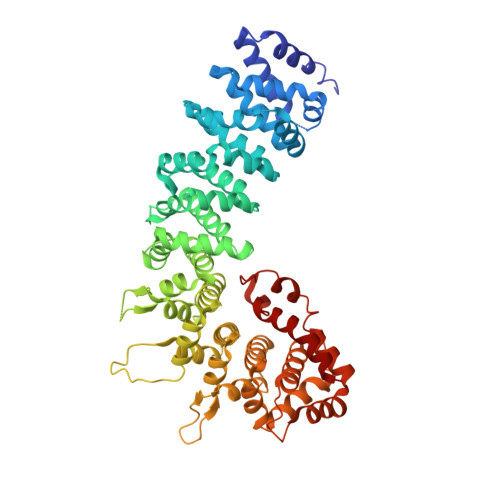
Unusual Armadillo Fold in the Human General Vesicular Transport Factor P115
Striegl, H., Roske, Y., Kummel, D., Heinemann, U.(2009) PLoS One 4: E4656
- PubMed: 19247479 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0004656
- Primary Citation Related Structures:
2W3C - PubMed Abstract:
The golgin family gives identity and structure to the Golgi apparatus and is part of a complex protein network at the Golgi membrane. The golgin p115 is targeted by the GTPase Rab1a, contains a large globular head region and a long region of coiled-coil which forms an extended rod-like structure. p115 serves as vesicle tethering factor and plays an important role at different steps of vesicular transport. Here we present the 2.2 A-resolution X-ray structure of the globular head region of p115. The structure exhibits an armadillo fold that is decorated by elongated loops and carries a C-terminal non-canonical repeat. This terminal repeat folds into the armadillo superhelical groove and allows homodimeric association with important implications for p115 mediated multiple protein interactions and tethering.
- Max-Delbrück-Centrum für Molekulare Medizin, Berlin, Germany.
Organizational Affiliation: