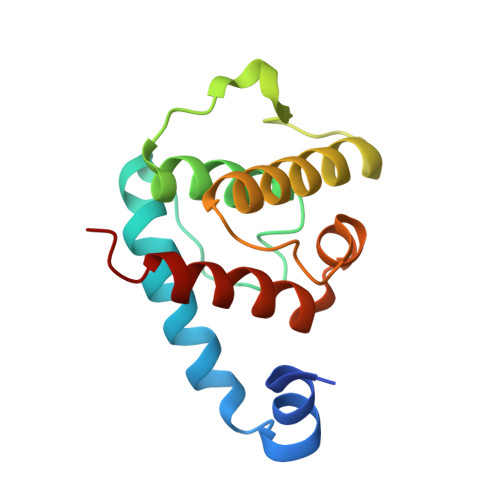
Structural analysis of the interactions between paxillin LD motifs and alpha-parvin.
Lorenz, S., Vakonakis, I., Lowe, E.D., Campbell, I.D., Noble, M.E., Hoellerer, M.K.(2008) Structure 16: 1521-1531
- PubMed: 18940607 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2008.08.007
- Primary Citation Related Structures:
2VZC, 2VZD, 2VZG, 2VZI - PubMed Abstract:
The adaptor protein paxillin contains five conserved leucine-rich (LD) motifs that interact with a variety of focal adhesion proteins, such as alpha-parvin. Here, we report the first crystal structure of the C-terminal calponin homology domain (CH(C)) of alpha-parvin at 1.05 A resolution and show that it is able to bind all the LD motifs, with some selectivity for LD1, LD2, and LD4. Cocrystal structures with these LD motifs reveal the molecular details of their interactions with a common binding site on alpha-parvin-CH(C), which is located at the rim of the canonical fold and includes part of the inter-CH domain linker. Surprisingly, this binding site can accommodate LD motifs in two antiparallel orientations. Taken together, these results reveal an unusual degree of binding degeneracy in the paxillin/alpha-parvin system that may facilitate the assembly of dynamic signaling complexes in the cell.
- Laboratory of Molecular Biophysics, University of Oxford, Oxford OX1 3QU, United Kingdom.
Organizational Affiliation: