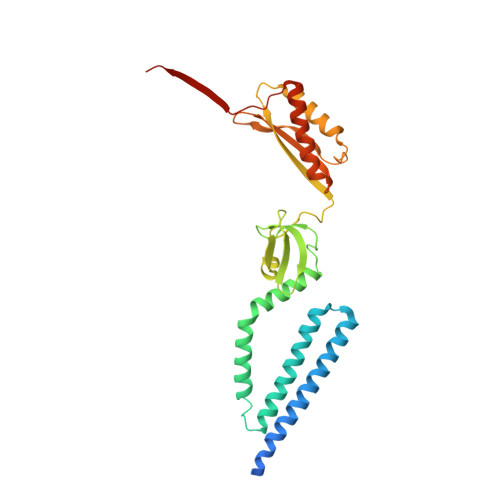
The Structure of an Open Form of an E. Coli Mechanosensitive Channel at 3.45 A Resolution.
Wang, W., Black, S.S., Edwards, M.D., Miller, S., Morrison, E.L., Bartlett, W., Dong, C., Naismith, J.H., Booth, I.R.(2008) Science 321: 1179
- PubMed: 18755969 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1159262
- Primary Citation Related Structures:
2VV5 - PubMed Abstract:
How ion channels are gated to regulate ion flux in and out of cells is the subject of intense interest. The Escherichia coli mechanosensitive channel, MscS, opens to allow rapid ion efflux, relieving the turgor pressure that would otherwise destroy the cell. We present a 3.45 angstrom-resolution structure for the MscS channel in an open conformation. This structure has a pore diameter of approximately 13 angstroms created by substantial rotational rearrangement of the three transmembrane helices. The structure suggests a molecular mechanism that underlies MscS gating and its decay of conductivity during prolonged activation. Support for this mechanism is provided by single-channel analysis of mutants with altered gating characteristics.
- Centre for Biomolecular Sciences, The North Haugh, University of St. Andrews, KY16 9ST, Scotland, UK.
Organizational Affiliation: