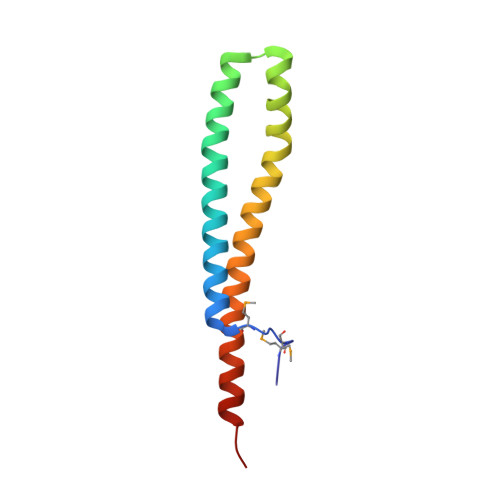
Structure of Staphylococcus Aureus Esxa Suggests a Contribution to Virulence by Action as a Transport Chaperone and/or Adaptor Protein.
Sundaramoorthy, R., Fyfe, P.K., Hunter, W.N.(2008) J Mol Biology 383: 603
- PubMed: 18773907 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.08.047
- Primary Citation Related Structures:
2VRZ, 2VS0 - PubMed Abstract:
Staphylococcus aureus pathogenesis depends on a specialized protein secretion system (ESX-1) that delivers a range of virulence factors to assist infectivity. We report the characterization of two such factors, EsxA and EsxB, small acidic dimeric proteins carrying a distinctive WXG motif. EsxA crystallized in triclinic and monoclinic forms and high-resolution structures were determined. The asymmetric unit of each crystal form is a dimer. The EsxA subunit forms an elongated cylindrical structure created from side-by-side alpha-helices linked with a hairpin bend formed by the WXG motif. Approximately 25% of the solvent accessible surface area of each subunit is involved in interactions, predominantly hydrophobic, with the partner subunit. Secondary-structure predictions suggest that EsxB displays a similar structure. The WXG motif helps to create a shallow cleft at each end of the dimer, forming a short beta-sheet-like feature with an N-terminal segment of the partner subunit. Structural and sequence comparisons, exploiting biological data on related proteins found in Mycobacterium tuberculosis, suggest that this family of proteins may contribute to pathogenesis by transporting protein cargo through the ESX-1 system exploiting a C-terminal secretion signal and/or are capable of acting as adaptor proteins to facilitate interactions with host receptor proteins.
- Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dundee, UK.
Organizational Affiliation: