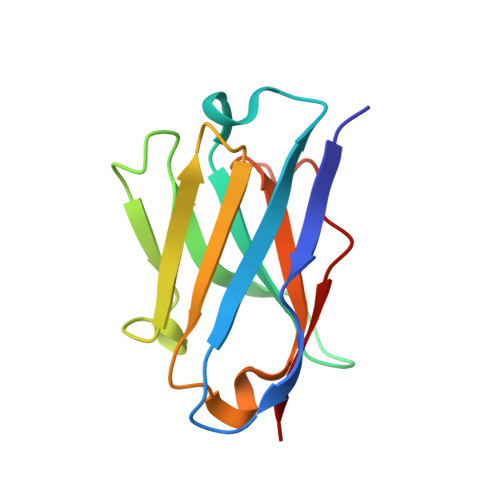
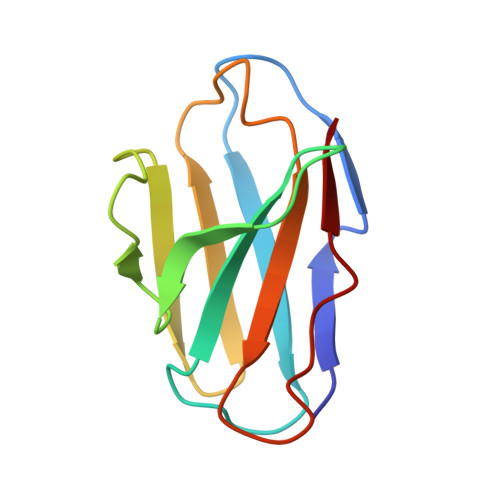
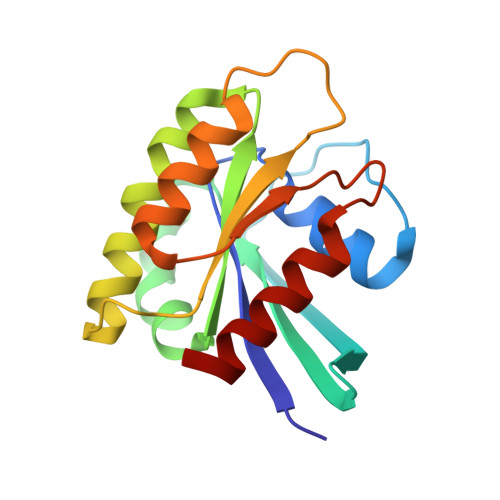
Functional Intracellular Antibody Fragments Do not Require Invariant Intra-Domain Disulfide Bonds.
Tanaka, T., Rabbitts, T.H.(2008) J Mol Biology 376: 749
- PubMed: 18187153 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.11.085
- Primary Citation Related Structures:
2VH5 - PubMed Abstract:
Intracellular antibody fragments that interfere with molecular interactions inside cells are valuable in investigation of interactomes and in therapeutics, but their application demands that they function in the reducing cellular milieu. We show here a 2.7-A crystal structure of intracellular antibody folds based on scaffolds developed from intracellular antibody capture technology, and we reveal that there is no structural or functional difference with or without the intra-domain disulfide bond of the variable domain of heavy chain or the variable domain of light chain. The data indicate that, in the reducing in vivo environment, the absence of the intra-domain disulfide bond is not an impediment to correction of antibody folding or to interaction with antigen. Thus, the structural constraints for in-cell function are intrinsic to variable single-domain framework sequences, providing a generic scaffold for isolation of functional intracellular antibody single domains.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 0QH, UK.
Organizational Affiliation: