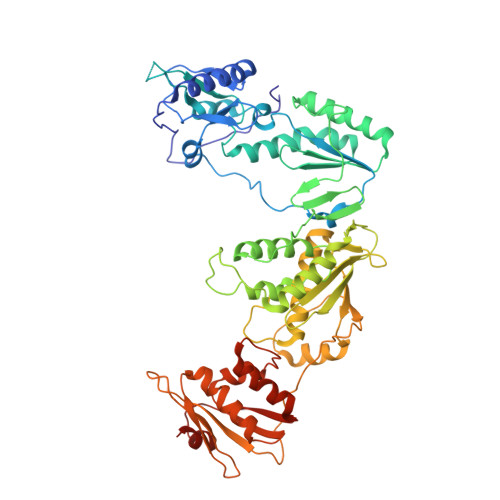
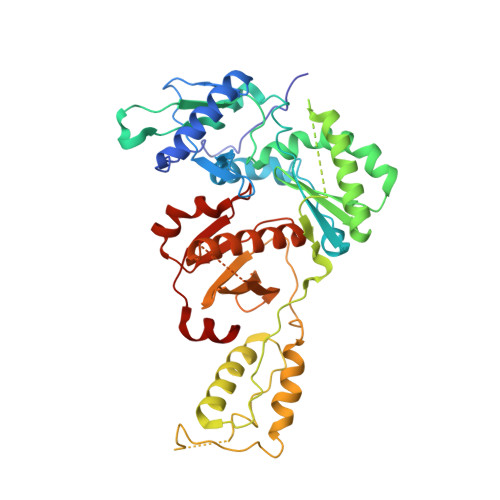
Crystal Structures of HIV-1 Reverse Transcriptase Complexes with Thiocarbamate Non-Nucleoside Inhibitors.
Spallarossa, A., Cesarini, S., Ranise, A., Ponassi, M., Unge, T., Bolognesi, M.(2008) Biochem Biophys Res Commun 365: 764
- PubMed: 18035053 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2007.11.036
- Primary Citation Related Structures:
2VG5, 2VG6, 2VG7 - PubMed Abstract:
O-Phthalimidoethyl-N-arylthiocarbamates (TCs) have been recently identified as a new class of potent HIV-1 reverse transcriptase (RT) non-nucleoside inhibitors (NNRTIs), by means of computer-aided drug design techniques [Ranise A. Spallarossa, S. Cesarini, F. Bondavalli, S. Schenone, O. Bruno, G. Menozzi, P. Fossa, L. Mosti, M. La Colla, et al., Structure-based design, parallel synthesis, structure-activity relationship, and molecular modeling studies of thiocarbamates, new potent non-nucleoside HIV-1 reverse transcriptase inhibitor isosteres of phenethylthiazolylthiourea derivatives, J. Med. Chem. 48 (2005) 3858-3873]. To elucidate the atomic details of RT/TC interaction and validate an earlier TC docking model, the structures of three RT/TC complexes were determined at 2.8-3.0A resolution by X-ray crystallography. The conformations adopted by the enzyme-bound TCs were analyzed and compared with those of bioisosterically related NNRTIs.
- Dipartimento di Scienze Farmaceutiche, Universita' di Genova, viale Benedetto XV, 3, I-16132 Genova, Italy.
Organizational Affiliation: