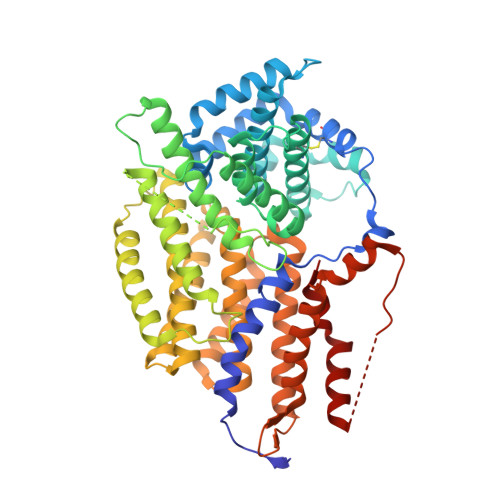
Insights Into Kinetochore-DNA Interactions from the Structure of Cep3Delta
Purvis, A., Singleton, M.R.(2008) EMBO Rep 9: 56
- PubMed: 18064045 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.embor.7401139
- Primary Citation Related Structures:
2VEQ - PubMed Abstract:
The CBF3 complex is an essential core component of the budding yeast kinetochore and is required for the centromeric localization of all other kinetochore proteins. We determined the crystal structure of a large section of the protein Cep3 from CBF3, which is the only component with obvious DNA-binding motifs. The protein adopts a roughly bilobal shape, with an extended dimerization interface. The dimer has a large central channel that is sufficient to accommodate duplex B-form DNA. The zinc-finger domains emerge at the edges of the channel, and could bind to the DNA in a pseudo-symmetrical manner at degenerate half-sites in the centromeric sequence. We propose a mechanism for the modulation of DNA affinity by an acidic activator domain, which could be applicable to a wider family of transcription factors.
- Macromolecular Structure and Function Laboratory, Cancer Research UK, London Research Institute, 44 Lincoln's Inn Fields, London, UK.
Organizational Affiliation: