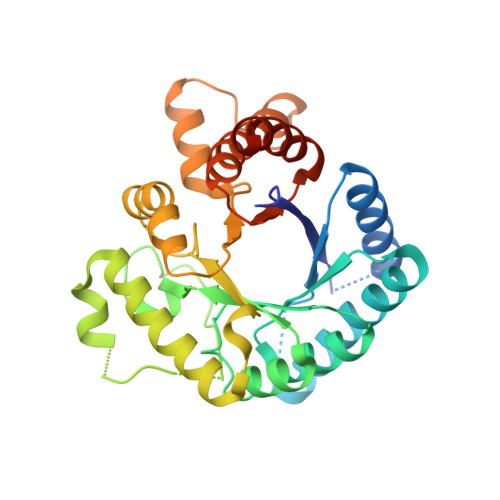
Dihydropteroate Synthase from Streptococcus Pneumoniae: Structure, Ligand Recognition and Mechanism of Sulfonamide Resistance.
Levy, C., Minnis, D., Derrick, J.P.(2008) Biochem J 412: 379
- PubMed: 18321242 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20071598
- Primary Citation Related Structures:
2VEF, 2VEG - PubMed Abstract:
DHPS (dihydropteroate synthase) catalyses an essential step in the biosynthesis of folic acid and is the target for the sulfonamide group of antimicrobial drugs. In the present paper we report two crystal structures of DHPS from the respiratory pathogen Streptococcus pneumoniae: the apoenzyme at 1.8 A (1 A=0.1 nm) resolution and a complex with DHPP (6-hydroxymethyl-7,8-dihydropterin monophosphate) at 2.4 A resolution. The enzyme forms a alpha/beta barrel structure, with a highly conserved binding pocket for recognition of the pterin substrate, DHPPP (6-hydroxymethyl-7,8-dihydropterin pyrophosphate). There is a fixed order of substrate binding: DHPPP binds first, followed by the second substrate, pABA (p-aminobenzoic acid). Binding of PP(i) also allows the enzyme to recognize pABA or sulfonamide drugs, which act as pABA analogues. Using equilibrium and pre-steady state kinetic fluorescence measurements, we show that the on-rate for DHPPP binding to the enzyme is relatively low (2.6x10(5) M(-1) x s(-1)) and propose that binding of this substrate induces a large scale movement of the second loop in the enzyme structure to participate in the formation of the pABA-binding site. Two mutations which confer resistance to sulfonamide drugs do not affect DHPPP binding, but have a substantial effect on pABA and sulfonamide recognition. The results show that binding of DHPPP and pABA are separate distinguishable events in the reaction cycle, and that mutations which confer resistance to sulfonamide drugs act exclusively on the second step in the binding process.
- Manchester Interdisciplinary Biocentre, University of Manchester, 131 Princess Street, Manchester M1 7DN, UK.
Organizational Affiliation: