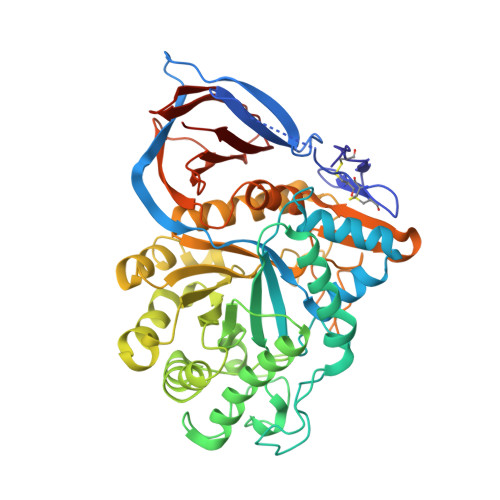
Crystal Structures of Complexes of N-Butyl- and N-Nonyl-Deoxynojirimycin Bound to Acid Beta-Glucosidase: Insights Into the Mechanism of Chemical Chaperone Action in Gaucher Disease.
Brumshtein, B., Greenblatt, H.M., Butters, T.D., Shaaltiel, Y., Aviezer, D., Silman, I., Futerman, A.H., Sussman, J.L.(2007) J Biological Chem 282: 29052
- PubMed: 17666401 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M705005200
- Primary Citation Related Structures:
2V3D, 2V3E - PubMed Abstract:
Gaucher disease is caused by mutations in the gene encoding acid beta-glucosidase (GlcCerase), resulting in glucosylceramide (GlcCer) accumulation. The only currently available orally administered treatment for Gaucher disease is N-butyl-deoxynojirimycin (Zavesca, NB-DNJ), which partially inhibits GlcCer synthesis, thus reducing levels of GlcCer accumulation. NB-DNJ also acts as a chemical chaperone for GlcCerase, although at a different concentration than that required to completely inhibit GlcCer synthesis. We now report the crystal structures, at 2A resolution, of complexes of NB-DNJ and N-nonyl-deoxynojirimycin (NN-DNJ) with recombinant human GlcCerase, expressed in cultured plant cells. Both inhibitors bind at the active site of GlcCerase, with the imino sugar moiety making hydrogen bonds to side chains of active site residues. The alkyl chains of NB-DNJ and NN-DNJ are oriented toward the entrance of the active site where they undergo hydrophobic interactions. Based on these structures, we make a number of predictions concerning (i) involvement of loops adjacent to the active site in the catalytic process, (ii) the nature of nucleophilic attack by Glu-340, and (iii) the role of a conserved water molecule located in a solvent cavity adjacent to the active site. Together, these results have significance for understanding the mechanism of action of GlcCerase and the mode of GlcCerase chaperoning by imino sugars.
- Departments of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: