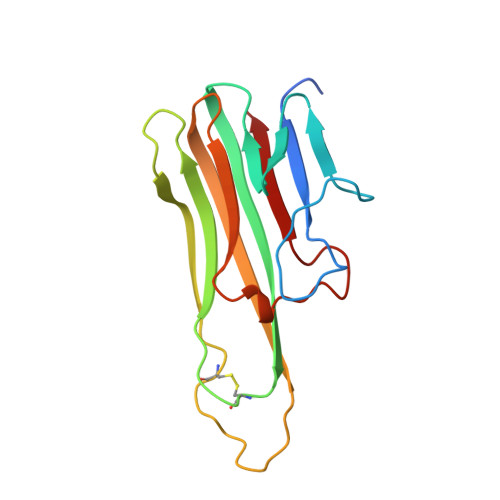
The structure of mouse tumour-necrosis factor at 1.4 A resolution: towards modulation of its selectivity and trimerization.
Baeyens, K.J., De Bondt, H.L., Raeymaekers, A., Fiers, W., De Ranter, C.J.(1999) Acta Crystallogr D Biol Crystallogr 55: 772-778
- PubMed: 10089307 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444998018435
- Primary Citation Related Structures:
2TNF - PubMed Abstract:
The 1.4 A resolution structure of recombinant mouse tumour-necrosis factor alpha (mTNF) at 100 K has been determined. The crystals are triclinic, space group P1, with unit-cell parameters a = 48.06, b = 48.18, c = 51.01 A, alpha = 114.8, beta = 103.6, gamma = 91.1 degrees. The structure was refined to a final crystallographic R value of 19.7% (Rfree = 23.3%), including 3477 protein atoms, one 2-propanol molecule, one Tris molecule and 240 water molecules. Throughout the crystal lattice, the trimers are differently packed compared with human TNF, which was crystallized in the tetragonal space group P41212 and refined to 2.6 A resolution. The structures of mTNF and human TNF are very similar, diverging mainly in regions that are either flexible and/or involved in crystal packing. Some loops in mTNF which contain residues important for receptor binding are better resolved than in human TNF, such as the surface-exposed loops 30-34 and 144-147, which are also important for receptor specificity. Compared with human TNFs, the channel formed by the three monomers in mTNF is narrower. One 2-propanol molecule trapped in the trimeric channel could be a lead compound for the design of TNF inhibitors.
- Laboratorium voor Analytische Chemie en Medicinale Fysicochemie, E. Van Evenstraat 4, B-3000 Leuven, Belgium.
Organizational Affiliation: