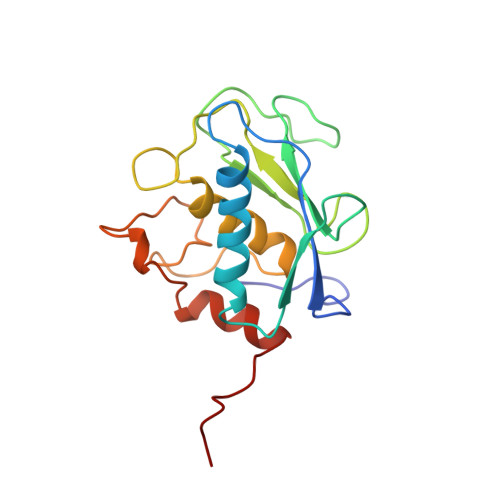
The NMR structure of the inhibited catalytic domain of human stromelysin-1.
Gooley, P.R., O'Connell, J.F., Marcy, A.I., Cuca, G.C., Salowe, S.P., Bush, B.L., Hermes, J.D., Esser, C.K., Hagmann, W.K., Springer, J.P., Johnson, B.A.(1994) Nat Struct Biol 1: 111-118
- PubMed: 7656014 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0294-111
- Primary Citation Related Structures:
2SRT - PubMed Abstract:
The three-dimensional structure of the catalytic domain of stromelysin-1 complexed with an N-carboxyl alkyl inhibitor has been determined by NMR methods. The global fold consists of three helices, a five stranded beta-sheet and a methionine located in a turn near the catalytic histidines, classifying stromelysin-1 as a metzincin. Stromelysin-1 is unique in having two independent zinc binding sites: a catalytic site and a structural site. The inhibitor binds in an extended conformation. The S1' subsite is a deep hydrophobic pocket, whereas S2' appears shallow and S3' open.
- Department of Biophysical Chemistry, Merck Research Laboratories, Rahway, New Jersey 07065-0900, USA.
Organizational Affiliation: