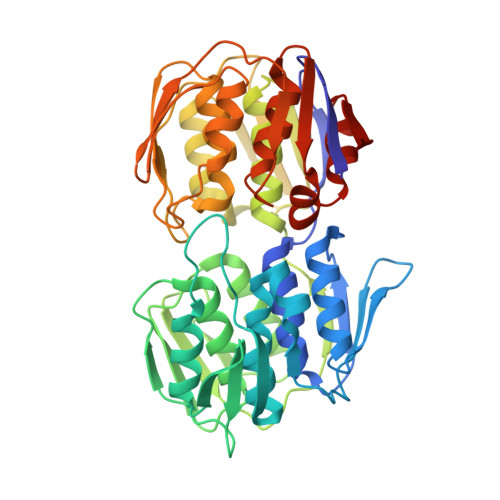
Crystal structure of UDP-N-acetylglucosamine enolpyruvyl transferase from Haemophilus influenzae in complex with UDP-N-acetylglucosamine and fosfomycin
Yoon, H.J., Lee, S.J., Mikami, B., Park, H.J., Yoo, J., Suh, S.W.(2008) Proteins 71: 1032-1037
- PubMed: 18247346 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21959
- Primary Citation Related Structures:
2RL1, 2RL2 - Department of Chemistry, College of Natural Sciences, Seoul National University, Seoul 151-747, Korea.
Organizational Affiliation: