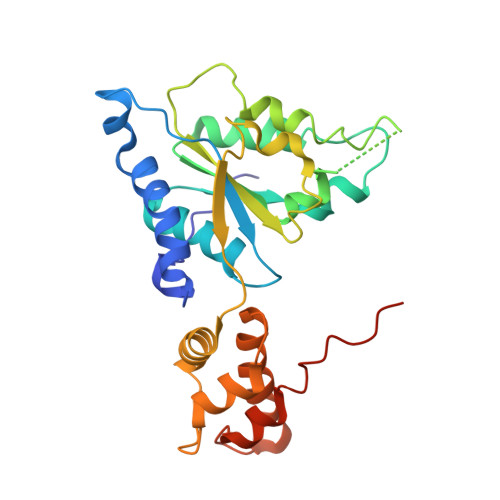
Structural studies on Helicobacter pyloriATP-dependent protease, FtsH
Kim, S.H., Kang, G.B., Song, H.-E., Park, S.J., Bea, M.-H., Eom, S.H.(2008) J Synchrotron Radiat 15: 208-210
- PubMed: 18421140 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S090904950706846X
- Primary Citation Related Structures:
2R62, 2R65 - PubMed Abstract:
The ATP-dependent protease, FtsH, degrades misassembled membrane proteins for quality control like SecY, subunit a of FoF1-ATPase, and YccA, and digests short-lived soluble proteins in order to control their cellular regulation, including sigma32, LpxC and lambdacII. The FtsH protein has an N-terminal transmembrane segment and a large cytosolic region that consists of two domains, an ATPase and a protease domain. To provide a structural basis for the nucleotide-dependent domain motions and a better understanding of substrate translocation, the crystal structures of the Helicobacter pylori (Hp) FtsH ATPase domain in the nucleotide-free state and complexed with ADP, were determined. Two different structures of HpFtsH ATPase were observed, with the nucleotide-free state in an asymmetric unit, and these structures reveal the new forms and show other conformational differences between the nucleotide-free and ADP-bound state compared with previous structures. In particular, one HpFtsH Apo structure has a considerable rotation difference compared with the HpFtsH ADP complex, and this large conformational change reveals that FtsH may have the mechanical force needed for substrate translocation.
- Department of Life Science, Cell Dynamics Research Center, Gwangju Institute of Science and Technology, Gwangju 500-712, Korea.
Organizational Affiliation: