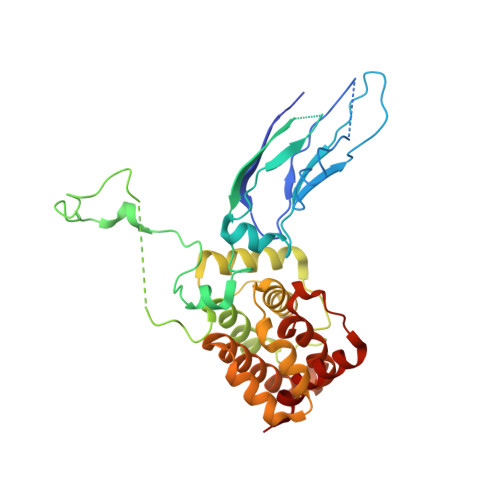
A role of the Lowe syndrome protein OCRL in early steps of the endocytic pathway
Erdmann, K.S., Mao, Y., McCrea, H.J., Zoncu, R., Lee, S., Paradise, S., Modregger, J., Biemesderfer, D., Toomre, D., De Camilli, P.(2007) Dev Cell 13: 377-390
- PubMed: 17765681 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.devcel.2007.08.004
- Primary Citation Related Structures:
2QV2 - PubMed Abstract:
Mutations in the inositol 5-phosphatase OCRL are responsible for Lowe syndrome, whose manifestations include mental retardation and renal Fanconi syndrome. OCRL has been implicated in membrane trafficking, but disease mechanisms remain unclear. We show that OCRL visits late-stage, endocytic clathrin-coated pits and binds the Rab5 effector APPL1 on peripheral early endosomes. The interaction with APPL1, which is mediated by the ASH-RhoGAP-like domains of OCRL and is abolished by disease mutations, provides a link to protein networks implicated in the reabsorptive function of the kidney and in the trafficking and signaling of growth factor receptors in the brain. Crystallographic studies reveal a role of the ASH-RhoGAP-like domains in positioning the phosphatase domain at the membrane interface and a clathrin box protruding from the RhoGAP-like domain. Our results support a role of OCRL in the early endocytic pathway, consistent with the predominant localization of its preferred substrates, PI(4,5)P(2) and PI(3,4,5)P(3), at the cell surface.
- Department of Cell Biology, School of Medicine, Yale University, New Haven, CT 06510, USA.
Organizational Affiliation: