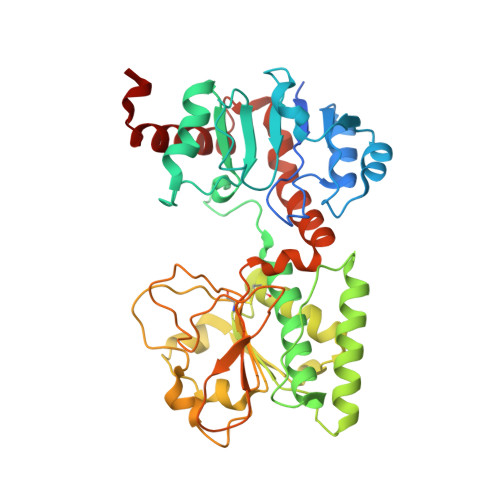
Crystal Structures of Ligand-Bound Saccharopine Dehydrogenase from Saccharomyces cerevisiae
Andi, B., Xu, H., Cook, P.F., West, A.H.(2007) Biochemistry 46: 12512-12521
- PubMed: 17939687 Search on PubMed
- DOI: https://doi.org/10.1021/bi701428m
- Primary Citation Related Structures:
2QRJ, 2QRK, 2QRL - PubMed Abstract:
Three structures of saccharopine dehydrogenase (l-lysine-forming) (SDH) have been determined in the presence of sulfate, adenosine monophosphate (AMP), and oxalylglycine (OxGly). In the sulfate-bound structure, a sulfate ion binds in a cleft between the two domains of SDH, occupies one of the substrate carboxylate binding sites, and results in partial closure of the active site of the enzyme due to a domain rotation of almost 12 degrees in comparison to the apoenzyme structure. In the second structure, AMP binds to the active site in an area where the NAD+ cofactor is expected to bind. All of the AMP moieties (adenine ring, ribose, and phosphate) interact with specific residues of the enzyme. In the OxGly-bound structure, carboxylates of OxGly interact with arginine residues representative of the manner in which substrate (alpha-ketoglutarate and saccharopine) may bind. The alpha-keto group of OxGly interacts with Lys77 and His96, which are candidates for acid-base catalysis. Analysis of ligand-enzyme interactions, comparative structural analysis, corroboration with kinetic data, and discussion of a ternary complex model are presented in this study.
- Department of Chemistry and Biochemistry, University of Oklahoma, 620 Parrington Oval, Norman, Oklahoma 73019, USA.
Organizational Affiliation: