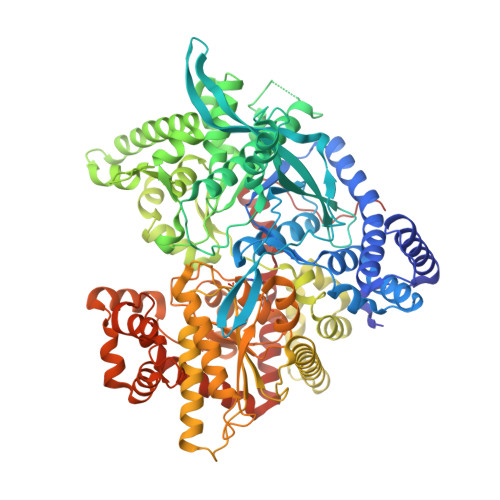
Naturally occurring pentacyclic triterpenes as inhibitors of glycogen phosphorylase: synthesis, structure-activity relationships, and X-ray crystallographic studies
Wen, X., Sun, H., Liu, J., Cheng, K., Zhang, P., Zhang, L., Hao, J., Zhang, L., Ni, P., Zographos, S.E., Leonidas, D.D., Alexacou, K.-M., Gimisis, T., Hayes, J.M., Oikonomakos, N.G.(2008) J Med Chem 51: 3540-3554
- PubMed: 18517260 Search on PubMed
- DOI: https://doi.org/10.1021/jm8000949
- Primary Citation Related Structures:
2QN1, 2QN2 - PubMed Abstract:
Twenty-five naturally occurring pentacyclic triterpenes, 15 of which were synthesized in this study, were biologically evaluated as inhibitors of rabbit muscle glycogen phosphorylase a (GPa). From SAR studies, the presence of a sugar moiety in triterpene saponins resulted in a markedly decreased activity ( 7, 18- 20) or no activity ( 21, 22). These saponins, however, might find their value as potential natural prodrugs which are much more water-soluble than their corresponding aglycones. To elucidate the mechanism of GP inhibition, we have determined the crystal structures of the GPb-asiatic acid and GPb-maslinic acid complexes. The X-ray analysis indicates that the inhibitors bind at the allosteric activator site, where the physiological activator AMP binds. Pentacyclic triterpenes represent a promising class of multiple-target antidiabetic agents that exert hypoglycemic effects, at least in part, through GP inhibition.
- Center for Drug Discovery, College of Pharmacy, China Pharmaceutical University, 24 Tongjia Xiang, Nanjing 210009, China.
Organizational Affiliation: