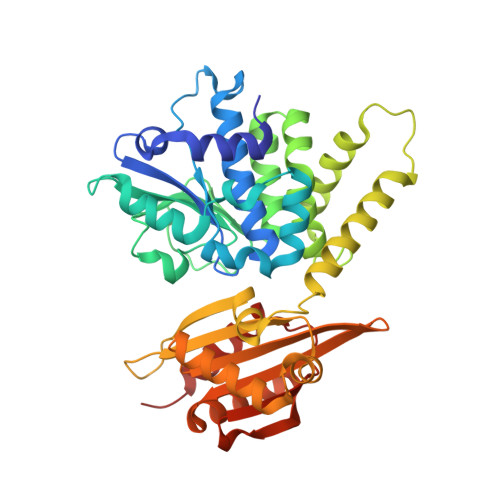
The crystal structure of the cytosolic exopolyphosphatase from Saccharomyces cerevisiae reveals the basis for substrate specificity.
Ugochukwu, E., Lovering, A.L., Mather, O.C., Young, T.W., White, S.A.(2007) J Mol Biology 371: 1007-1021
- PubMed: 17599355 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.05.066
- Primary Citation Related Structures:
2QB6, 2QB7, 2QB8 - PubMed Abstract:
Inorganic long-chain polyphosphate is a ubiquitous linear polymer in biology, consisting of many phosphate moieties linked by phosphoanhydride bonds. It is synthesized by polyphosphate kinase, and metabolised by a number of enzymes, including exo- and endopolyphosphatases. The Saccharomyces cerevisiae gene PPX1 encodes for a 45 kDa, metal-dependent, cytosolic exopolyphosphatase that processively cleaves the terminal phosphate group from the polyphosphate chain, until inorganic pyrophosphate is all that remains. PPX1 belongs to the DHH family of phosphoesterases, which includes: family-2 inorganic pyrophosphatases, found in Gram-positive bacteria; prune, a cyclic AMPase; and RecJ, a single-stranded DNA exonuclease. We describe the high-resolution X-ray structures of yeast PPX1, solved using the multiple isomorphous replacement with anomalous scattering (MIRAS) technique, and its complexes with phosphate (1.6 A), sulphate (1.8 A) and ATP (1.9 A). Yeast PPX1 folds into two domains, and the structures reveal a strong similarity to the family-2 inorganic pyrophosphatases, particularly in the active-site region. A large, extended channel formed at the interface of the N and C-terminal domains is lined with positively charged amino acids and represents a conduit for polyphosphate and the site of phosphate hydrolysis. Structural comparisons with the inorganic pyrophosphatases and analysis of the ligand-bound complexes lead us to propose a hydrolysis mechanism. Finally, we discuss a structural basis for substrate selectivity and processivity.
- The School of Biosciences, The University of Birmingham, Edgbaston, Birmingham, B15 2TT, UK.
Organizational Affiliation: