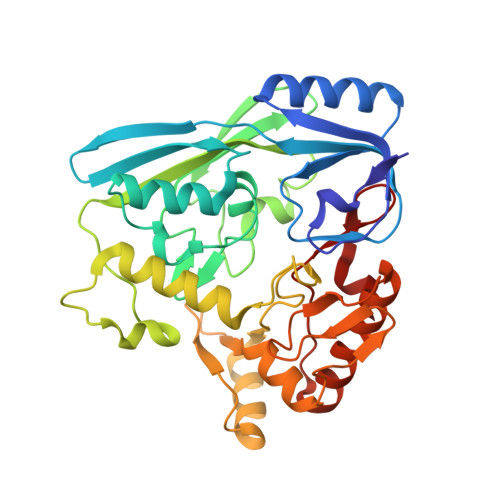
Crystal Structure of E. Coli Mur B bound to a napthyl tetronic acid inhibitor
Mansour, T., Caulfield, C.E., Rasmussen, B., Chopra, R., Krishnamurthy, G., Morris, K.M., Svenson, K., Bard, J., Smeltzer, C., Naughton, S., Antane, S., Yang, Y., Severin, A., Quagliato, D., Petersen, P.J., Singh, G.To be published.