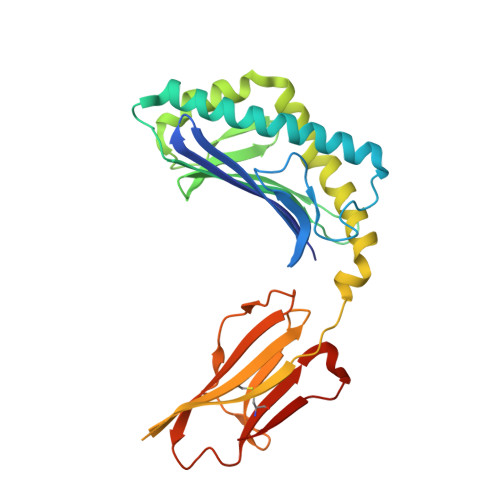
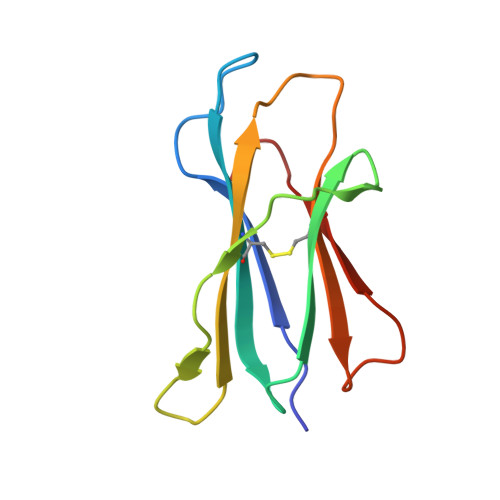
Crystal Structures of Mouse CD1d-iGb3 Complex and its Cognate Valpha14 T Cell Receptor Suggest a Model for Dual Recognition of Foreign and Self Glycolipids.
Zajonc, D.M., Savage, P.B., Bendelac, A., Wilson, I.A., Teyton, L.(2008) J Mol Biology 377: 1104-1116
- PubMed: 18295796 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.01.061
- Primary Citation Related Structures:
2Q7Y, 2Q86 - PubMed Abstract:
The semi-invariant Valpha14Jalpha18 T cell receptor (TCR) is expressed by regulatory NKT cells and has the unique ability to recognize chemically diverse ligands presented by CD1d. The crystal structure of CD1d complexed to a natural, endogenous ligand, isoglobotrihexosylceramide (iGb3), illustrates the extent of this diversity when compared to the binding of potent, exogenous ligands, such as alpha-galactosylceramide (alpha-GalCer). A single mode of recognition for these two classes of ligands would then appear problematic for a single T cell receptor. However, the Valpha14 TCR adopts two different conformations in the crystal where, in one configuration, the presence of a larger cavity between the two CDR3 regions could accommodate iGb3 and, in the other, a smaller cavity fits alpha-GalCer more snugly. Alternatively, the extended iGb3 headgroup could be "squashed" upon docking of the TCR and accommodated between the CD1 and TCR surfaces. Thus, the same TCR may adopt alternative modes of recognition for these foreign and self-ligands for NKT cell activation.
- Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Rd., La Jolla, CA 92037, USA.
Organizational Affiliation: