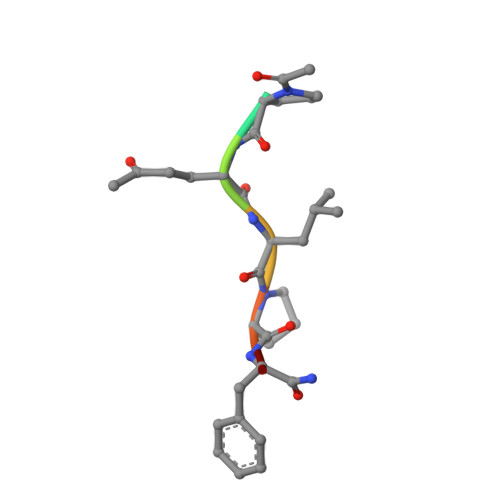
Transglutaminase 2 undergoes a large conformational change upon activation
Pinkas, D.M., Strop, P., Brunger, A.T., Khosla, C.(2007) PLoS Biol 5: e327-e327
- PubMed: 18092889 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.0050327
- Primary Citation Related Structures:
2Q3Z - PubMed Abstract:
Human transglutaminase 2 (TG2), a member of a large family of enzymes that catalyze protein crosslinking, plays an important role in the extracellular matrix biology of many tissues and is implicated in the gluten-induced pathogenesis of celiac sprue. Although vertebrate transglutaminases have been studied extensively, thus far all structurally characterized members of this family have been crystallized in conformations with inaccessible active sites. We have trapped human TG2 in complex with an inhibitor that mimics inflammatory gluten peptide substrates and have solved, at 2-A resolution, its x-ray crystal structure. The inhibitor stabilizes TG2 in an extended conformation that is dramatically different from earlier transglutaminase structures. The active site is exposed, revealing that catalysis takes place in a tunnel, bridged by two tryptophan residues that separate acyl-donor from acyl-acceptor and stabilize the tetrahedral reaction intermediates. Site-directed mutagenesis was used to investigate the acyl-acceptor side of the tunnel, yielding mutants with a marked increase in preference for hydrolysis over transamidation. By providing the ability to visualize this activated conformer, our results create a foundation for understanding the catalytic as well as the non-catalytic roles of TG2 in biology, and for dissecting the process by which the autoantibody response to TG2 is induced in celiac sprue patients.
- Department of Chemical Engineering, Stanford University, Stanford, California, United States of America.
Organizational Affiliation: