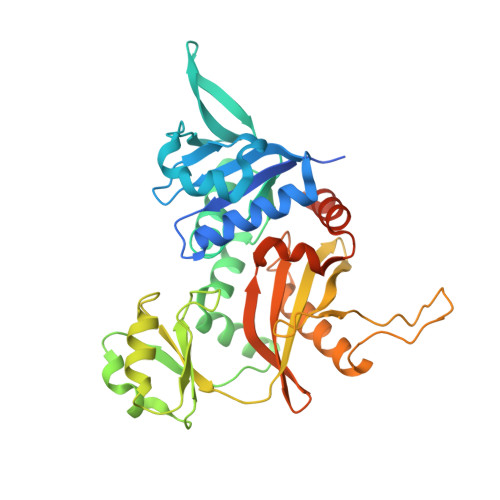
Enzymatic characterization and crystal structure analysis of the D-alanine-D-alanine ligase from Helicobacter pylori.
Wu, D., Zhang, L., Kong, Y., Du, J., Chen, S., Chen, J., Ding, J., Jiang, H., Shen, X.(2008) Proteins 72: 1148-1160
- PubMed: 18320587 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22009
- Primary Citation Related Structures:
2PVP - PubMed Abstract:
D-Alanine-D-alanine ligase is the second enzyme in the D-Ala branch of bacterial cell wall peptidoglycan assembly, and recognized as an attractive antimicrobial target. In this work, the D-Ala-D-Ala ligase of Helicobacter pylori strain SS1 (HpDdl) was kinetically and structurally characterized. The determined apparent K(m) of ATP (0.87 microM), the K(m1) (1.89 mM) and K(m2) of D-Ala (627 mM), and the k(cat) (115 min(-1)) at pH 8.0 indicated its relatively weak binding affinity and poor catalytic activity against the substrate D-Ala in vitro. However, by complementary assay of expressing HpDdl in Escherichia coli Delta ddl mutant, HpDdl was confirmed to be capable of D-Ala-D-Ala ligating in vivo. Through sequence alignment with other members of the D-Ala-D-X ligase superfamily, HpDdl keeps two conservatively substituted residues (Ile16 and Leu241) and two nonconserved residues (Leu308 and Tyr311) broadly located in the active region of the enzyme. Kinetic analyses against the corresponding HpDdl mutants (I16V, L241Y, L241F, L308T, and Y311S) suggested that these residues, especially Leu308 and Tyr311, might partly contribute to the unique catalytic properties of the enzyme. This was fairly proved by the crystal structure of HpDdl, which revealed that there is a 3(10)-helix (including residues from Gly306 to Leu312) near the D-Ala binding region in the C-terminal domain, where HpDdl has two sequence deletions compared with other homologs. Such 3(10)-helix may participate in D-Ala binding and conformational change of the enzyme. Our present work hopefully provides useful information for understanding the D-Ala-D-Ala ligase of Helicobacter pylori.
- Drug Discovery and Design Center, State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai 201203, China.
Organizational Affiliation: