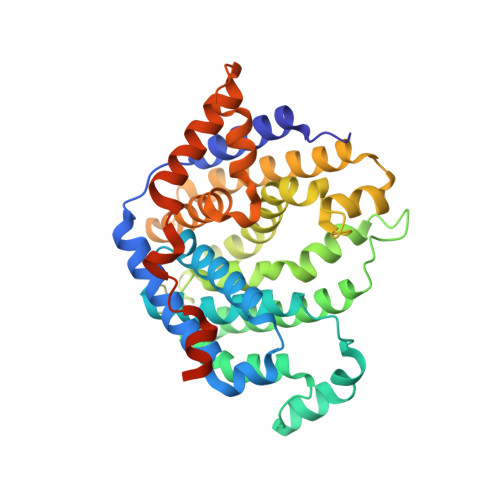
Structural and mechanistic analysis of trichodiene synthase using site-directed mutagenesis: probing the catalytic function of tyrosine-295 and the asparagine-225/serine-229/glutamate-233-Mg2+B motif.
Vedula, L.S., Jiang, J., Zakharian, T., Cane, D.E., Christianson, D.W.(2008) Arch Biochem Biophys 469: 184-194
- PubMed: 17996718 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.abb.2007.10.015
- Primary Citation Related Structures:
2PS4, 2PS5, 2PS6, 2PS7, 2PS8 - PubMed Abstract:
Trichodiene synthase from Fusarium sporotrichioides contains two metal ion-binding motifs required for the cyclization of farnesyl diphosphate: the "aspartate-rich" motif D(100)DXX(D/E) that coordinates to Mg2+A and Mg2+C, and the "NSE/DTE" motif N(225)DXXSXXXE that chelates Mg2+B (boldface indicates metal ion ligands). Here, we report steady-state kinetic parameters, product array analyses, and X-ray crystal structures of trichodiene synthase mutants in which the fungal NSE motif is progressively converted into a plant-like DDXXTXXXE motif, resulting in a degradation in both steady-state kinetic parameters and product specificity. Each catalytically active mutant generates a different distribution of sesquiterpene products, and three newly detected sesquiterpenes are identified. In addition, the kinetic and structural properties of the Y295F mutant of trichodiene synthase were found to be similar to those of the wild-type enzyme, thereby ruling out a proposed role for Y295 in catalysis.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, PA 19104-6323, USA.
Organizational Affiliation: