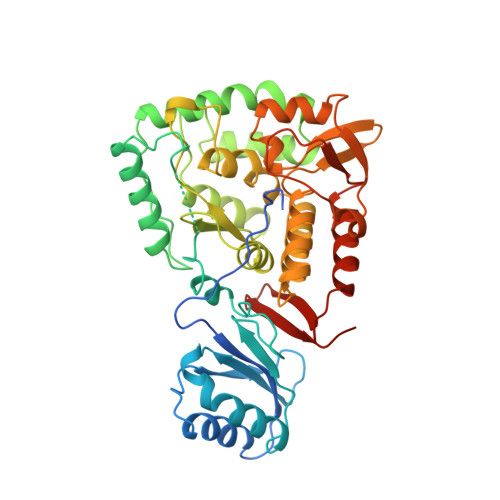
Structural basis of autoregulation of phenylalanine hydroxylase.
Kobe, B., Jennings, I.G., House, C.M., Michell, B.J., Goodwill, K.E., Santarsiero, B.D., Stevens, R.C., Cotton, R.G., Kemp, B.E.(1999) Nat Struct Biol 6: 442-448
- PubMed: 10331871 Search on PubMed
- DOI: https://doi.org/10.1038/8247
- Primary Citation Related Structures:
1PHZ, 2PHM - PubMed Abstract:
Phenylalanine hydroxylase converts phenylalanine to tyrosine, a rate-limiting step in phenylalanine catabolism and protein and neurotransmitter biosynthesis. It is tightly regulated by the substrates phenylalanine and tetrahydrobiopterin and by phosphorylation. We present the crystal structures of dephosphorylated and phosphorylated forms of a dimeric enzyme with catalytic and regulatory properties of the wild-type protein. The structures reveal a catalytic domain flexibly linked to a regulatory domain. The latter consists of an N-terminal autoregulatory sequence (containing Ser 16, which is the site of phosphorylation) that extends over the active site pocket, and an alpha-beta sandwich core that is, unexpectedly, structurally related to both pterin dehydratase and the regulatory domains of metabolic enzymes. Phosphorylation has no major structural effects in the absence of phenylalanine, suggesting that phenylalanine and phosphorylation act in concert to activate the enzyme through a combination of intrasteric and possibly allosteric mechanisms.
- St. Vincent's Institute of Medical Research, Fitzroy, Victoria, Australia. B.Kobe@medicine.unimelb.edu.au
Organizational Affiliation: