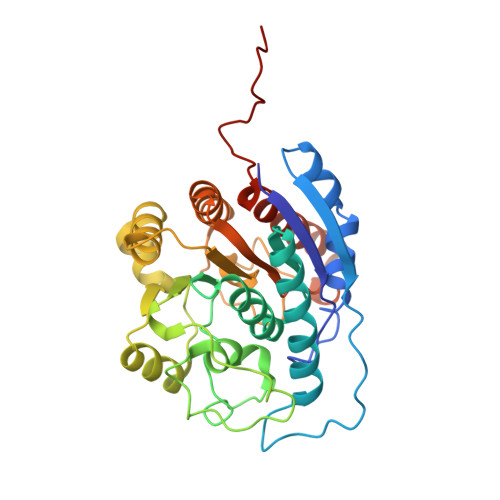
Crystal structure of human arginase I complexed with thiosemicarbazide reveals an unusual thiocarbonyl mu-sulfide ligand in the binuclear manganese cluster.
Di Costanzo, L., Pique, M.E., Christianson, D.W.(2007) J Am Chem Soc 129: 6388-6389
- PubMed: 17469833 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja071567j
- Primary Citation Related Structures:
2PHA, 2PHO, 2ZAV - PubMed Abstract:
The crystal structure of the human arginase I-thiosemicarbazide complex reveals an unusual thiocarbonyl μ-sulfide ligand in the binuclear manganese cluster. The C=S moiety of thiosemicarbazide bridges Mn 2+ A and Mn 2+ B with coordination distances of 2.6 Å and 2.4 Å, respectively. Otherwise, the binding of thiosemicarbazide to human arginase I does not cause any significant structural changes in the active site. The crystal structure of the unliganded enzyme reveals a hydrogen bonded water molecule that could support proton transfer between a μ-water molecule and H141 to regenerate the nucleophilic μ-hydroxide ion in the final step of catalysis.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, PA 19104-6323, USA.
Organizational Affiliation: