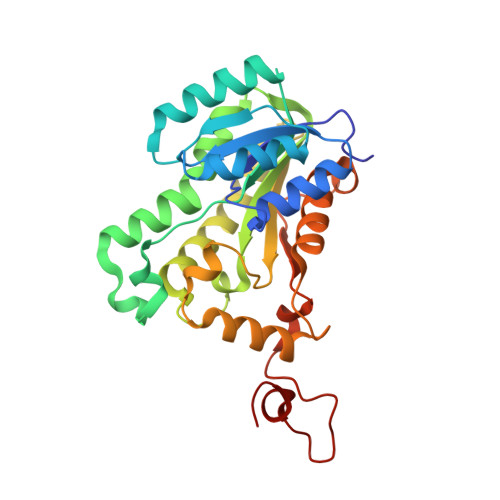
Crystal structure of the Helicobacter pylori enoyl-acyl carrier protein reductase in complex with hydroxydiphenyl ether compounds, triclosan and diclosan
Lee, H.H., Moon, J.H., Suh, S.W.(2007) Proteins 69: 691-694
- PubMed: 17879346 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21586
- Primary Citation Related Structures:
2PD3, 2PD4 - Department of Chemistry, College of Natural Sciences, Seoul National University, Seoul 151-742, Korea.
Organizational Affiliation: