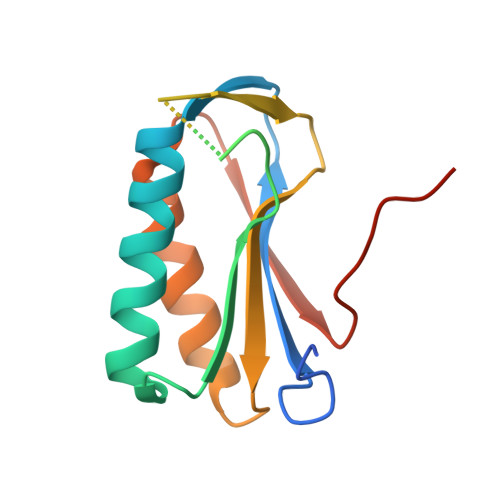
Crystal Structure of Human Ribosomal Protein L10 Core Domain Reveals Eukaryote-Specific Motifs in Addition to the Conserved Fold
Nishimura, M., Kaminishi, T., Takemoto, C., Kawazoe, M., Yoshida, T., Tanaka, A., Sugano, S., Shirouzu, M., Ohkubo, T., Yokoyama, S., Kobayashi, Y.(2008) J Mol Biology 377: 421-430
- PubMed: 18258260 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.01.003
- Primary Citation Related Structures:
2PA2 - PubMed Abstract:
A phylogenetically conserved ribosomal protein L16p/L10e organizes the architecture of the aminoacyl tRNA binding site on the large ribosomal subunit. Eukaryotic L10 also exhibits a variety of cellular activities, and, in particular, human L10 is known as a putative tumor suppressor, QM. We have determined the 2.5-A crystal structure of the human L10 core domain that corresponds to residues 34-182 of the full-length 214 amino acids. Its two-layered alpha+beta architecture is significantly similar to those of the archaeal and bacterial homologues, substantiating a high degree of structural conservation across the three phylogenetic domains. A cation-binding pocket formed between alpha2 and beta 6 is similar to that of the archaeal L10 protein but appears to be better ordered. Previously reported L10 mutations that cause defects in the yeast ribosome are clustered around this pocket, indicating that its integrity is crucial for its role in L10 function. Characteristic interactions among Arg90-Trp171-Arg139 guide the C-terminal part outside of the central fold, implying that the eukaryote-specific C-terminal extension localizes on the outer side of the ribosome.
- Graduate School of Pharmaceutical Sciences, Osaka University, 1-6 Yamadaoka, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: