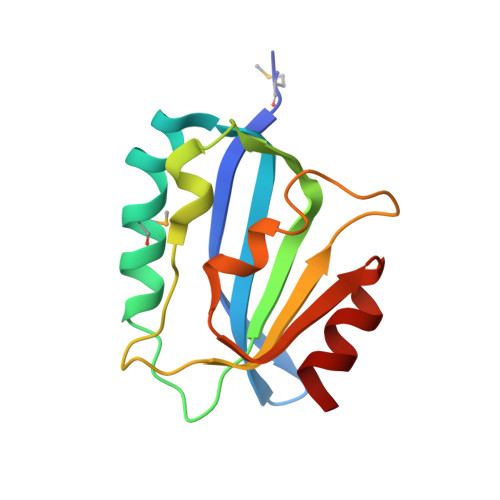
Open and closed conformations of two SpoIIAA-like proteins (YP_749275.1 and YP_001095227.1) provide insights into membrane association and ligand binding.
Kumar, A., Lomize, A., Jin, K.K., Carlton, D., Miller, M.D., Jaroszewski, L., Abdubek, P., Astakhova, T., Axelrod, H.L., Chiu, H.J., Clayton, T., Das, D., Deller, M.C., Duan, L., Feuerhelm, J., Grant, J.C., Grzechnik, A., Han, G.W., Klock, H.E., Knuth, M.W., Kozbial, P., Krishna, S.S., Marciano, D., McMullan, D., Morse, A.T., Nigoghossian, E., Okach, L., Reyes, R., Rife, C.L., Sefcovic, N., Tien, H.J., Trame, C.B., van den Bedem, H., Weekes, D., Xu, Q., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1245-1253
- PubMed: 20944218 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109042481
- Primary Citation Related Structures:
2OOK, 2Q3L - PubMed Abstract:
The crystal structures of the proteins encoded by the YP_749275.1 and YP_001095227.1 genes from Shewanella frigidimarina and S. loihica, respectively, have been determined at 1.8 and 2.25 Å resolution, respectively. These proteins are members of a novel family of bacterial proteins that adopt the α/β SpoIIAA-like fold found in STAS and CRAL-TRIO domains. Despite sharing 54% sequence identity, these two proteins adopt distinct conformations arising from different dispositions of their α2 and α3 helices. In the `open' conformation (YP_001095227.1), these helices are 15 Å apart, leading to the creation of a deep nonpolar cavity. In the `closed' structure (YP_749275.1), the helices partially unfold and rearrange, occluding the cavity and decreasing the solvent-exposed hydrophobic surface. These two complementary structures are reminiscent of the conformational switch in CRAL-TRIO carriers of hydrophobic compounds. It is suggested that both proteins may associate with the lipid bilayer in their `open' monomeric state by inserting their amphiphilic helices, α2 and α3, into the lipid bilayer. These bacterial proteins may function as carriers of nonpolar substances or as interfacially activated enzymes.
- Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.
Organizational Affiliation: